showhmmprof
Plot hidden Markov model (HMM) profile
Syntax
showhmmprof(Model)
showhmmprof(Model,
...'Scale', ScaleValue, ...)
showhmmprof(Model, ...'Order', OrderValue,
...)
Arguments
Model | Hidden Markov model created by the function gethmmprof or pfamhmmread. |
ScaleValue | Property to select a probability scale. Enter one of the following
values:
|
OrderValue | Property to specify the order of the amino acid alphabet. Enter a character vector or string
with the 20 standard amino acids characters A R N D C Q E G H I
L K M F P S T W Y V. The ambiguous characters B Z
X are not allowed. |
Description
showhmmprof( plots
a profile hidden Markov model described by the structure Model)Model.
showhmmprof(..., ' calls PropertyName', PropertyValue,
...)showhmmprof with optional
properties that use property name/property value pairs. You can specify
one or more properties in any order. Each PropertyName must
be enclosed in single quotation marks and is case insensitive. These
property name/property value pairs are as follows:
showhmmprof(specifies
the scale to use. If log probabilities (Model,
...'Scale', ScaleValue, ...) ScaleValue='logprob'ScaleValue='prob'ScaleValue='logodds'ScaleValue'logprob'.
showhmmprof( specifies the order in which the symbols are arranged
along the vertical axis. This option allows you reorder the alphabet
and group the symbols according to their properties.Model, ...'Order', OrderValue,
...)
Examples
Load a model example.
model = pfamhmmread('pf00002.ls');Plot the profile.
showhmmprof(model, 'Scale', 'logodds')


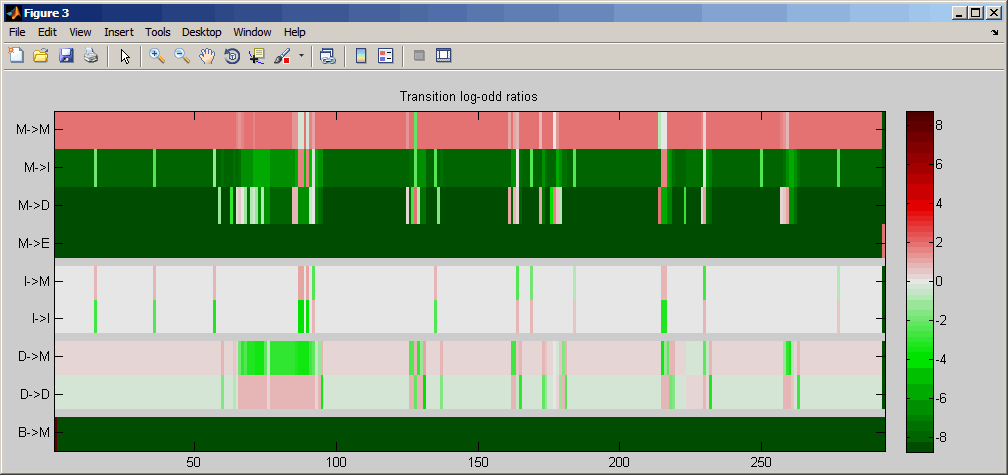
Order the alphabet by hydrophobicity.
hydrophobic = 'IVLFCMAGTSWYPHNDQEKR';Plot the profile.
showhmmprof(model, 'Order', hydrophobic)


Version History
Introduced before R2006a
See Also
gethmmprof | hmmprofalign | hmmprofestimate | hmmprofgenerate | hmmprofstruct | pfamhmmread
