imhmin
Suppress regional minima in image using H-minima transform
Description
J = imhmin(I,H)I by using the
H-minima transform. The H-minima transform decreases the depth of all regional
minima by an amount up to H. As a result, the transform fully
suppresses regional minima whose depth is less than H.
Regional minima are connected pixels with the same
intensity value, t, that are surrounded by pixels with an
intensity value greater than t.
Examples
Calculate H-Minima Transform
Create a 10-by-10 sample image. Add three regional minima, each consisting of an area of connected pixels surrounded by higher intensity values.
a = 10*ones(10,10); a(2:4,2:4) = 7; a(6:8,6:8) = 2; a(1:3,7:9) = 13; a(2,8) = 10;
This image is a grayscale representation of the pixel values. The depth of each minimum depends on the surrounding pixel values.

Apply the H-minima transform that decreases the depth of regional minima by up to 4.
h = 4; b = imhmin(a,h);
This image is a grayscale representation of the transformed image. The transform fully suppresses two of the minima. The transform partially suppresses the deepest minimum, and adds 4 to the intensity values of the pixels in that minimum.

Improve Watershed Segmentation Using H-Minima Transform
You can suppress shallow regional minima to avoid oversegmentation during watershed segmentation.
Load an RGB image of pears to segment. Convert the image to grayscale and display it. The center of each pear is bright, corresponding to a regional maximum.
RGB = imread("pears.png");
I = im2gray(RGB);
imshow(I)
In watershed segmentation, the image is analogous to a surface comprised of watershed lines and catchment basins. When water flows into the surface, it pools in the catchment basins. In a grayscale image, local minima are the catchment basins. To segment the pears, invert the image so the centers of the pears become regional minima.
Icomp = imcomplement(I); imshow(Icomp)

Display the inverted image as a 3-D surface, in which the third dimension for each pixel is its intensity value. The deeper regions for each pear have spiky bottoms, indicating many shallow regional minima, like catchment basins into which water can pool.
surf(Icomp,EdgeColor="none")
colormap(gray)
Segment the unfiltered image and display the result as a label overlay. The image is oversegmented, meaning there are many small masks rather than one mask for each pear.
L = watershed(Icomp); overlay = labeloverlay(I,L); imshow(overlay)

Suppress the shallow minima by applying the H-minima transform. The value for h has been determined by using trial and error. Change the value to see how the h value affects the segmentation result.
h =  30;
Ifilt = imhmin(Icomp,h);
30;
Ifilt = imhmin(Icomp,h);Display the filtered image as a 3-D surface.
surf(Ifilt,EdgeColor="none")
colormap(gray)
Segment the filtered image and display the result. The image contains approximately one mask for each pear in the foreground.
Lfilt = watershed(Ifilt); overlayfilt = labeloverlay(I,Lfilt); imshow(overlayfilt)

Input Arguments
I — Input image
numeric array
Input image, specified as a numeric array of any dimension.
Data Types: single | double | int8 | int16 | int32 | int64 | uint8 | uint16 | uint32 | uint64
H — H-minima transform
nonnegative scalar
H-minima transform, specified as a nonnegative scalar.
Data Types: single | double | int8 | int16 | int32 | int64 | uint8 | uint16 | uint32 | uint64
conn — Pixel connectivity
4 | 8 | 6 | 18 | 26 | 3-by-3-by- ... -by-3 matrix of 0s and
1s
Pixel connectivity, specified as one of the values in this table. The
default connectivity is 8 for 2-D images, and
26 for 3-D images.
Value | Meaning | |
|---|---|---|
Two-Dimensional Connectivities | ||
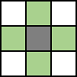
| Pixels are connected if their edges touch. The neighborhood of a pixel are the adjacent pixels in the horizontal or vertical direction. |
Current pixel is shown in gray. |
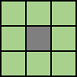
| Pixels are connected if their edges or corners touch. The neighborhood of a pixel are the adjacent pixels in the horizontal, vertical, or diagonal direction. |
Current pixel is shown in gray. |
Three-Dimensional Connectivities | ||
| Pixels are connected if their faces touch. The neighborhood of a pixel are the adjacent pixels in:
|
Current pixel is shown in gray. |
| Pixels are connected if their faces or edges touch. The neighborhood of a pixel are the adjacent pixels in:
|
Current pixel is center of cube. |
| Pixels are connected if their faces, edges, or corners touch. The neighborhood of a pixel are the adjacent pixels in:
|
Current pixel is center of cube. |
For higher dimensions, imhmin uses the default value
conndef(ndims(I),"maximal")
Connectivity can also be
defined in a more general way for any dimension by specifying a 3-by-3-by- ... -by-3 matrix of
0s and 1s. The 1-valued elements
define neighborhood locations relative to the center element of conn. Note
that conn must be symmetric about its center element. See Specifying Custom Connectivities for more information.
Data Types: single | double | int8 | int16 | int32 | int64 | uint8 | uint16 | uint32 | uint64
Output Arguments
J — Transformed image
numeric array
Transformed image, returned as a numeric array of the same size and data
type as I.
References
[1] Soille, P. Morphological Image Analysis: Principles and Applications. Springer-Verlag, 1999, pp. 170-171.
Extended Capabilities
C/C++ Code Generation
Generate C and C++ code using MATLAB® Coder™.
Usage notes and limitations:
imhminsupports the generation of C code (requires MATLAB® Coder™). Note that if you choose the genericMATLAB Host Computertarget platform,imhmingenerates code that uses a precompiled, platform-specific shared library. Use of a shared library preserves performance optimizations but limits the target platforms for which code can be generated. For more information, see Types of Code Generation Support in Image Processing Toolbox.When generating code, the optional third input argument,
conn, must be a compile-time constant.
Version History
Introduced before R2006a
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.

Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list:
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)