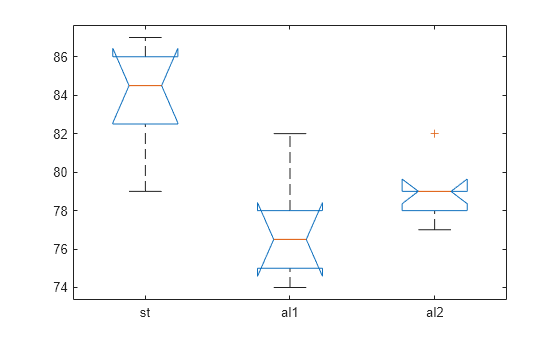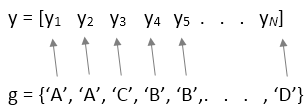anova1
One-way analysis of variance
Syntax
Description
p = anova1(y)y and returns the p-value.
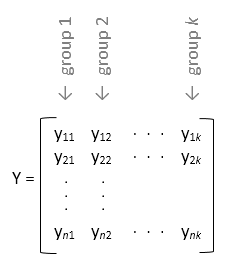
anova1 treats each column of y as a
separate group. The function tests the hypothesis that the samples in the columns of
y are drawn from populations with the same mean against the
alternative hypothesis that the population means are not all the same. The function
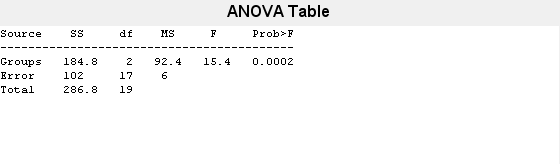
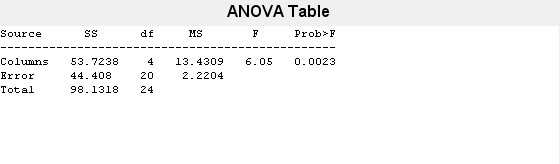
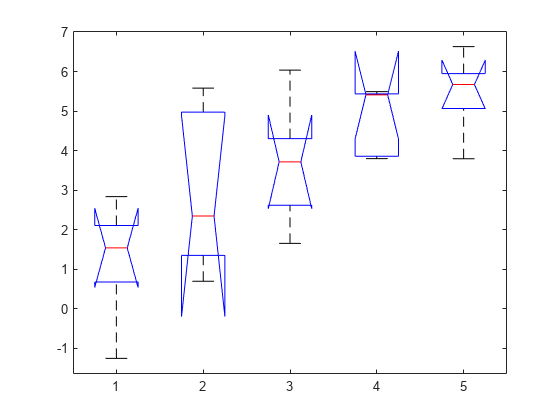
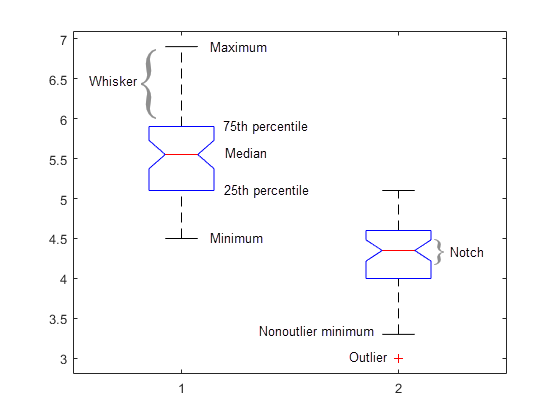
also displays the box plot for each group
in y and the standard ANOVA table
(tbl).
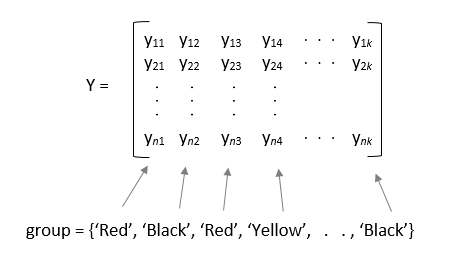
p = anova1(y,group,displayopt)displayopt is 'on' (default)
and suppresses the displays when displayopt is 'off'.
[ returns a structure, p,tbl,stats]
= anova1(___)stats,
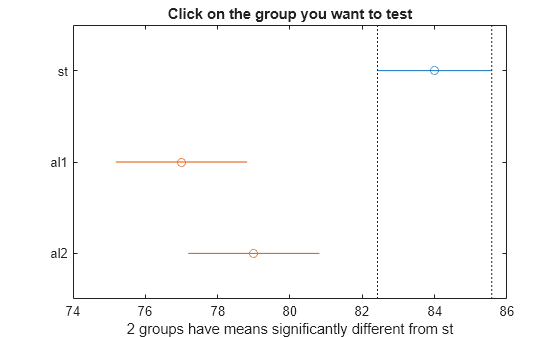
which you can use to perform a multiple comparison test. A multiple
comparison test enables you to determine which pairs of group means
are significantly different. To perform this test, use multcompare, providing the stats structure
as an input argument.
Examples
Input Arguments
Output Arguments
More About
Alternative Functionality
Instead of using anova1, you can create an anova
object by using the anova function.
The anova function provides these advantages:
The
anovafunction allows you to specify the ANOVA model type, sum of squares type, and factors to treat as categorical.anovaalso supports table predictor and response input arguments.In addition to the outputs returned by
anova1, the properties of theanovaobject contain the following:ANOVA model formula
Fitted ANOVA model coefficients
Residuals
Factors and response data
The
anovaobject functions allow you to conduct further analysis after fitting theanovaobject. For example, you can create an interactive plot of multiple comparisons of means for the ANOVA, get the mean response estimates for each value of a factor, and calculate the variance component estimates.
References
[1] Hogg, R. V., and J. Ledolter. Engineering Statistics. New York: MacMillan, 1987.
Version History
Introduced before R2006a
See Also
anova | anova2 | anovan | boxplot | multcompare