Decision Trees
Decision trees, or classification trees and regression trees, predict responses to
data. To predict a response, follow the decisions in the tree from the root (beginning)
node down to a leaf node. The leaf node contains the response. Classification trees give
responses that are nominal, such as 'true' or
'false'. Regression trees give numeric responses.
Statistics and Machine Learning Toolbox™ trees are binary. Each step in a prediction involves checking the value of one predictor (variable). For example, here is a simple classification tree:
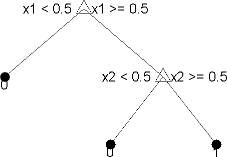
This tree predicts classifications based on two predictors, x1 and
x2. To predict, start at the top node, represented by a triangle
(Δ). The first decision is whether x1 is smaller than
0.5. If so, follow the left branch, and see that the tree
classifies the data as type 0.
If, however, x1 exceeds 0.5, then follow the
right branch to the lower-right triangle node. Here the tree asks if
x2 is smaller than 0.5. If so, then follow the
left branch to see that the tree classifies the data as type 0. If
not, then follow the right branch to see that the tree classifies the data as type
1.
To learn how to prepare your data for classification or regression using decision trees, see Steps in Supervised Learning.
Train Classification Tree
This example shows how to train a classification tree.
Create a classification tree using the entire ionosphere data set.
load ionosphere % Contains X and Y variables Mdl = fitctree(X,Y)
Mdl =
ClassificationTree
ResponseName: 'Y'
CategoricalPredictors: []
ClassNames: {'b' 'g'}
ScoreTransform: 'none'
NumObservations: 351
Train Regression Tree
This example shows how to train a regression tree.
Create a regression tree using all observation in the carsmall data set. Consider the Horsepower and Weight vectors as predictor variables, and the MPG vector as the response.
load carsmall % Contains Horsepower, Weight, MPG X = [Horsepower Weight]; Mdl = fitrtree(X,MPG)
Mdl =
RegressionTree
ResponseName: 'Y'
CategoricalPredictors: []
ResponseTransform: 'none'
NumObservations: 94
References
[1] Breiman, L., J. H. Friedman, R. A. Olshen, and C. J. Stone. Classification and Regression Trees. Boca Raton, FL: Chapman & Hall, 1984.
See Also
fitctree | fitrtree | ClassificationTree | RegressionTree
