Hidden Markov Models (HMM)
Introduction to Hidden Markov Models (HMM)
A hidden Markov model (HMM) is one in which you observe a sequence of emissions, but do not know the sequence of states the model went through to generate the emissions. Analyses of hidden Markov models seek to recover the sequence of states from the observed data.
As an example, consider a Markov model with two states and six possible emissions. The model uses:
A red die, having six sides, labeled 1 through 6.
A green die, having twelve sides, five of which are labeled 2 through 6, while the remaining seven sides are labeled 1.
A weighted red coin, for which the probability of heads is .9 and the probability of tails is .1.
A weighted green coin, for which the probability of heads is .95 and the probability of tails is .05.
The model creates a sequence of numbers from the set {1, 2, 3, 4, 5, 6} with the following rules:
Begin by rolling the red die and writing down the number that comes up, which is the emission.
Toss the red coin and do one of the following:
If the result is heads, roll the red die and write down the result.
If the result is tails, roll the green die and write down the result.
At each subsequent step, you flip the coin that has the same color as the die you rolled in the previous step. If the coin comes up heads, roll the same die as in the previous step. If the coin comes up tails, switch to the other die.
The state diagram for this model has two states, as shown in the following figure.
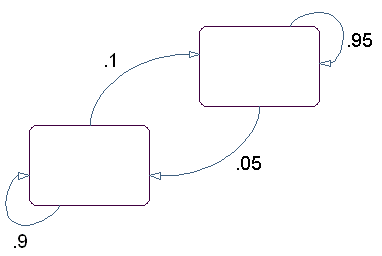
You determine the emission from a state by rolling the die with the same color as the state. You determine the transition to the next state by flipping the coin with the same color as the state.
The transition matrix is:
The emissions matrix is:
The model is not hidden because you know the sequence of states from the colors of the coins and dice. Suppose, however, that someone else is generating the emissions without showing you the dice or the coins. All you see is the sequence of emissions. If you start seeing more 1s than other numbers, you might suspect that the model is in the green state, but you cannot be sure because you cannot see the color of the die being rolled.
Hidden Markov models raise the following questions:
Given a sequence of emissions, what is the most likely state path?
Given a sequence of emissions, how can you estimate transition and emission probabilities of the model?
What is the forward probability that the model generates a given sequence?
What is the posterior probability that the model is in a particular state at any point in the sequence?
Analyzing Hidden Markov Models
Statistics and Machine Learning Toolbox™ functions related to hidden Markov models are:
hmmgenerate— Generates a sequence of states and emissions from a Markov modelhmmestimate— Calculates maximum likelihood estimates of transition and emission probabilities from a sequence of emissions and a known sequence of stateshmmtrain— Calculates maximum likelihood estimates of transition and emission probabilities from a sequence of emissionshmmviterbi— Calculates the most probable state path for a hidden Markov modelhmmdecode— Calculates the posterior state probabilities of a sequence of emissions
This section shows how to use these functions to analyze hidden Markov models.
Generating a Test Sequence
The following commands create the transition and emission matrices for the model described in the Introduction to Hidden Markov Models (HMM):
TRANS = [.9 .1; .05 .95]; EMIS = [1/6, 1/6, 1/6, 1/6, 1/6, 1/6;... 7/12, 1/12, 1/12, 1/12, 1/12, 1/12];
To generate a random sequence of states and emissions from the
model, use hmmgenerate:
[seq,states] = hmmgenerate(1000,TRANS,EMIS);
The output seq is the sequence of emissions
and the output states is the sequence of states.
hmmgenerate begins in state 1 at step 0,
makes the transition to state i1 at
step 1, and returns i1 as
the first entry in states. To change the initial
state, see Changing the Initial State Distribution.
Estimating the State Sequence
Given the transition and emission matrices TRANS and EMIS,
the function hmmviterbi uses
the Viterbi algorithm to compute the most likely sequence of states
the model would go through to generate a given sequence seq of
emissions:
likelystates = hmmviterbi(seq, TRANS, EMIS);
likelystates is a sequence the same length
as seq.
To test the accuracy of hmmviterbi, compute
the percentage of the actual sequence states that
agrees with the sequence likelystates.
sum(states==likelystates)/1000 ans = 0.8200
In this case, the most likely sequence of states agrees with the random sequence 82% of the time.
Estimating Transition and Emission Matrices
The functions hmmestimate and hmmtrain estimate the transition and emission
matrices TRANS and EMIS given
a sequence seq of emissions.
Using hmmestimate. The function hmmestimate requires that you
know the sequence of states states that the model went through to generate seq.
The following takes the emission and state sequences and returns estimates of the transition and emission matrices:
[TRANS_EST, EMIS_EST] = hmmestimate(seq, states) TRANS_EST = 0.8989 0.1011 0.0585 0.9415 EMIS_EST = 0.1721 0.1721 0.1749 0.1612 0.1803 0.1393 0.5836 0.0741 0.0804 0.0789 0.0726 0.1104
You can compare the outputs with the original transition and emission matrices,
TRANS and EMIS:
TRANS TRANS = 0.9000 0.1000 0.0500 0.9500 EMIS EMIS = 0.1667 0.1667 0.1667 0.1667 0.1667 0.1667 0.5833 0.0833 0.0833 0.0833 0.0833 0.0833
Using hmmtrain. If you do not know the sequence of states states, but you
have initial guesses for TRANS and EMIS, you
can still estimate TRANS and EMIS using
hmmtrain.
Suppose you have the following initial guesses for TRANS and
EMIS.
TRANS_GUESS = [.85 .15; .1 .9]; EMIS_GUESS = [.17 .16 .17 .16 .17 .17;.6 .08 .08 .08 .08 08];
You estimate TRANS and EMIS as
follows:
[TRANS_EST2, EMIS_EST2] = hmmtrain(seq, TRANS_GUESS, EMIS_GUESS) TRANS_EST2 = 0.2286 0.7714 0.0032 0.9968 EMIS_EST2 = 0.1436 0.2348 0.1837 0.1963 0.2350 0.0066 0.4355 0.1089 0.1144 0.1082 0.1109 0.1220
hmmtrain uses an iterative algorithm that alters the
matrices TRANS_GUESS and EMIS_GUESS so that
at each step the adjusted matrices are more likely to generate the observed
sequence, seq. The algorithm halts when the matrices in two
successive iterations are within a small tolerance of each other.
If the algorithm fails to reach this tolerance within a maximum number of
iterations, whose default value is 100, the algorithm halts. In
this case, hmmtrain returns the last values of
TRANS_EST and EMIS_EST and issues a
warning that the tolerance was not reached.
If the algorithm fails to reach the desired tolerance, increase the default value of the maximum number of iterations with the command:
hmmtrain(seq,TRANS_GUESS,EMIS_GUESS,'maxiterations',maxiter)
where maxiter is the maximum number of steps the algorithm
executes.
Change the default value of the tolerance with the command:
hmmtrain(seq, TRANS_GUESS, EMIS_GUESS, 'tolerance', tol)
where tol is the desired value of the tolerance. Increasing
the value of tol makes the algorithm halt sooner, but the
results are less accurate.
Two factors reduce the reliability of the output matrices of
hmmtrain:
The algorithm converges to a local maximum that does not represent the true transition and emission matrices. If you suspect this, use different initial guesses for the matrices
TRANS_ESTandEMIS_EST.The sequence
seqmay be too short to properly train the matrices. If you suspect this, use a longer sequence forseq.
Estimating Posterior State Probabilities
The posterior state probabilities of an emission sequence seq are
the conditional probabilities that the model is in a particular state
when it generates a symbol in seq, given that seq is
emitted. You compute the posterior state probabilities with hmmdecode:
PSTATES = hmmdecode(seq,TRANS,EMIS)
The output PSTATES is an M-by-L matrix,
where M is the number of states and L is
the length of seq. PSTATES(i,j) is
the conditional probability that the model is in state i when
it generates the jth symbol of seq,
given that seq is emitted.
hmmdecode begins with the model in state
1 at step 0, prior to the first emission. PSTATES(i,1) is
the probability that the model is in state i at the following step
1. To change the initial state, see Changing the Initial State Distribution.
To return the logarithm of the probability of the sequence seq,
use the second output argument of hmmdecode:
[PSTATES,logpseq] = hmmdecode(seq,TRANS,EMIS)
The probability of a sequence tends to 0 as the length of the
sequence increases, and the probability of a sufficiently long sequence
becomes less than the smallest positive number your computer can represent. hmmdecode returns
the logarithm of the probability to avoid this problem.
Changing the Initial State Distribution
By default, Statistics and Machine Learning Toolbox hidden Markov model functions begin in state 1. In other words, the distribution of initial states has all of its probability mass concentrated at state 1. To assign a different distribution of probabilities, p = [p1, p2, ..., pM], to the M initial states, do the following:
Create an M+1-by-M+1 augmented transition matrix, of the following form:
where T is the true transition matrix. The first column of contains M+1 zeros. p must sum to 1.
Create an M+1-by-N augmented emission matrix, , that has the following form:
If the transition and emission matrices are TRANS and EMIS,
respectively, you create the augmented matrices with the following
commands:
TRANS_HAT = [0 p; zeros(size(TRANS,1),1) TRANS]; EMIS_HAT = [zeros(1,size(EMIS,2)); EMIS];
See Also
hmmdecode | hmmestimate | hmmgenerate | hmmtrain | hmmviterbi
