k-Means Clustering
This topic provides an introduction to k-means clustering and an
example that uses the Statistics and Machine Learning Toolbox™ function kmeans to find the best clustering solution
for a data set.
Introduction to k-Means Clustering
k-means clustering is a partitioning method. The function kmeans partitions data into k mutually exclusive
clusters and returns the index of the cluster to which it assigns each observation.
kmeans treats each observation in your data as an object
that has a location in space. The function finds a partition in which objects within
each cluster are as close to each other as possible, and as far from objects in
other clusters as possible. You can choose a distance metric to use with
kmeans based on attributes of your data. Like many
clustering methods, k-means clustering requires you to specify
the number of clusters k before clustering.
Unlike hierarchical clustering, k-means clustering operates on actual observations rather than the dissimilarity between every pair of observations in the data. Also, k-means clustering creates a single level of clusters, rather than a multilevel hierarchy of clusters. Therefore, k-means clustering is often more suitable than hierarchical clustering for large amounts of data.
Each cluster in a k-means partition consists of member objects and a
centroid (or center). In each cluster, kmeans minimizes the sum
of the distances between the centroid and all member objects of the cluster.
kmeans computes centroid clusters differently for the
supported distance metrics. For details, see 'Distance'.
You can control the details of the minimization using name-value pair arguments available to
kmeans; for example, you can specify the initial values of
the cluster centroids and the maximum number of iterations for the algorithm. By
default, kmeans uses the k-means++ algorithm to initialize cluster
centroids, and the squared Euclidean distance metric to determine distances.
When performing k-means clustering, follow these best practices:
Compare k-means clustering solutions for different values of k to determine an optimal number of clusters for your data.
Evaluate clustering solutions by examining silhouette plots and silhouette values. You can also use the
evalclustersfunction to evaluate clustering solutions based on criteria such as gap values, silhouette values, Davies-Bouldin index values, and Calinski-Harabasz index values.Replicate clustering from different randomly selected centroids and return the solution with the lowest total sum of distances among all the replicates.
To perform k-means clustering interactively, use the Cluster Data Live Editor task.
Compare k-Means Clustering Solutions
This example explores k-means clustering on a four-dimensional data set. The example shows how to determine the correct number of clusters for the data set by using silhouette plots and values to analyze the results of different k-means clustering solutions. The example also shows how to use the 'Replicates' name-value pair argument to test a specified number of possible solutions and return the one with the lowest total sum of distances.
Load Data Set
Load the kmeansdata data set.
rng('default') % For reproducibility load('kmeansdata.mat') size(X)
ans = 1×2
560 4
The data set is four-dimensional and cannot be visualized easily. However, kmeans enables you to investigate whether a group structure exists in the data.
Create Clusters and Examine Separation
Partition the data set into three clusters using k-means clustering. Specify the city block distance metric, and use the default k-means++ algorithm for cluster center initialization. Use the 'Display' name-value pair argument to print the final sum of distances for the solution.
[idx3,C,sumdist3] = kmeans(X,3,'Distance','cityblock', ... 'Display','final');
Replicate 1, 7 iterations, total sum of distances = 2459.98. Best total sum of distances = 2459.98
idx3 contains cluster indices that indicate the cluster assignment for each row in X. To see if the resulting clusters are well separated, you can create a silhouette plot.
A silhouette plot displays a measure of how close each point in one cluster is to points in the neighboring clusters. This measure ranges from 1 (indicating points that are very distant from neighboring clusters) through 0 (points that are not distinctly in one cluster or another) to –1 (points that are probably assigned to the wrong cluster). silhouette returns these values in its first output.
Create a silhouette plot from idx3. Specify 'cityblock' for the distance metric to indicate that the k-means clustering is based on the sum of absolute differences.
[silh3,h] = silhouette(X,idx3,'cityblock'); xlabel('Silhouette Value') ylabel('Cluster')

The silhouette plot shows that most points in the second cluster have a large silhouette value (greater than 0.6), indicating that the cluster is somewhat separated from neighboring clusters. However, the third cluster contains many points with low silhouette values, and the first and third clusters contain a few points with negative values, indicating that these two clusters are not well separated.
To see if kmeans can find a better grouping of the data, increase the number of clusters to four. Print information about each iteration by using the 'Display' name-value pair argument.
idx4 = kmeans(X,4,'Distance','cityblock','Display','iter');
iter phase num sum
1 1 560 1792.72
2 1 6 1771.1
Best total sum of distances = 1771.1
Create a silhouette plot for the four clusters.
[silh4,h] = silhouette(X,idx4,'cityblock'); xlabel('Silhouette Value') ylabel('Cluster')
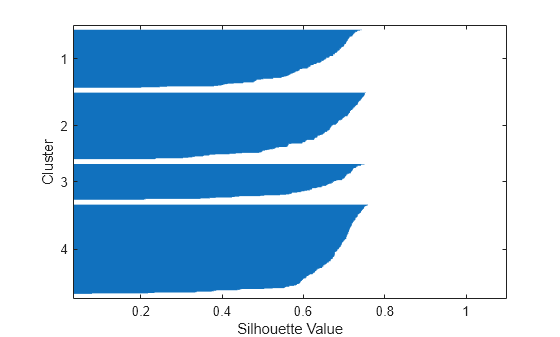
The silhouette plot indicates that these four clusters are better separated than the three clusters in the previous solution. You can take a more quantitative approach to comparing the two solutions by computing the average silhouette values for the two cases.
Compute the average silhouette values.
cluster3 = mean(silh3)
cluster3 = 0.5352
cluster4 = mean(silh4)
cluster4 = 0.6400
The average silhouette value of the four clusters is higher than the average value of the three clusters. These values support the conclusion represented in the silhouette plots.
Finally, find five clusters in the data. Create a silhouette plot and compute the average silhouette values for the five clusters.
idx5 = kmeans(X,5,'Distance','cityblock','Display','final');
Replicate 1, 7 iterations, total sum of distances = 1647.26. Best total sum of distances = 1647.26
[silh5,h] = silhouette(X,idx5,'cityblock'); xlabel('Silhouette Value') ylabel('Cluster')

mean(silh5)
ans = 0.5721
The silhouette plot indicates that five is probably not the right number of clusters, because two clusters contain points with mostly low silhouette values, and the fifth cluster contains a few points with negative values. Also, the average silhouette value for the five clusters is lower than the value for the four clusters. Without knowing how many clusters are in the data, it is a good idea to experiment with a range of values for k, the number of clusters.
Note that the sum of distances decreases as the number of clusters increases. For example, the sum of distances decreases from 2459.98 to 1771.1 to 1647.26 as the number of clusters increases from 3 to 4 to 5. Therefore, the sum of distances is not useful for determining the optimal number of clusters.
Avoid Local Minima
By default, kmeans begins the clustering process using a randomly selected set of initial centroid locations. The kmeans algorithm can converge to a solution that is a local (nonglobal) minimum; that is, kmeans can partition the data such that moving any single point to a different cluster increases the total sum of distances. However, as with many other types of numerical minimizations, the solution that kmeans reaches sometimes depends on the starting points. Therefore, other solutions (local minima) that have lower total sums of distances can exist for the data. You can use the 'Replicates' name-value pair argument to test different solutions. When you specify more than one replicate, kmeans repeats the clustering process starting from different randomly selected centroids for each replicate, and returns the solution with the lowest total sum of distances among all the replicates.
Find four clusters in the data and replicate the clustering five times. Also, specify the city block distance metric, and use the 'Display' name-value pair argument to print the final sum of distances for each solution.
[idx4,cent4,sumdist] = kmeans(X,4,'Distance','cityblock', ... 'Display','final','Replicates',5);
Replicate 1, 2 iterations, total sum of distances = 1771.1. Replicate 2, 3 iterations, total sum of distances = 1771.1. Replicate 3, 3 iterations, total sum of distances = 1771.1. Replicate 4, 6 iterations, total sum of distances = 2300.23. Replicate 5, 2 iterations, total sum of distances = 1771.1. Best total sum of distances = 1771.1
In replicate 4, kmeans finds a local minimum. Because each replicate begins from a different randomly selected set of initial centroids, kmeans sometimes finds more than one local minimum. However, the final solution that kmeans returns is the one with the lowest total sum of distances over all replicates.
Find the total of the within-cluster sums of point-to-centroid distances for the final solution returned by kmeans.
sum(sumdist)
ans = 1.7711e+03
See Also
Cluster
Data | kmeans | silhouette
