predict
Predict response of linear mixed-effects model
Syntax
Description
ypred = predict(lme)ypred at the original predictors
used to fit the linear mixed-effects model lme.
ypred = predict(lme,tblnew)ypred from
the fitted linear mixed-effects model lme at the
values in the new table or dataset array tblnew.
Use a table or dataset array for predict if you
use a table or dataset array for fitting the model lme.
If a particular grouping variable in tblnew has
levels that are not in the original data, then the random effects
for that grouping variable do not contribute to the 'Conditional' prediction
at observations where the grouping variable has new levels.
ypred = predict(lme,Xnew,Znew)ypred from
the fitted linear mixed-effects model lme at the
values in the new fixed- and random-effects design matrices, Xnew and Znew,
respectively. Znew can also be a cell array of
matrices. In this case, the grouping variable G is ones(n,1),
where n is the number of observations used in the
fit.
Use the matrix format for predict if using
design matrices for fitting the model lme.
ypred = predict(lme,Xnew,Znew,Gnew)ypred from
the fitted linear mixed-effects model lme at the
values in the new fixed- and random-effects design matrices, Xnew and Znew,
respectively, and the grouping variable Gnew.
Znew and Gnew can also
be cell arrays of matrices and grouping variables, respectively.
ypred = predict(___,Name,Value)ypred from the
fitted linear mixed-effects model lme with additional
options specified by one or more Name,Value pair
arguments.
For example, you can specify the confidence level, simultaneous confidence bounds, or contributions from only fixed effects.
Examples
Predict Responses at the Original Design Values
Load the sample data.
load('fertilizer.mat');The dataset array includes data from a split-plot experiment, where soil is divided into three blocks based on the soil type: sandy, silty, and loamy. Each block is divided into five plots, where five different types of tomato plants (cherry, heirloom, grape, vine, and plum) are randomly assigned to these plots. The tomato plants in the plots are then divided into subplots, where each subplot is treated by one of four fertilizers. This is simulated data.
Store the data in a dataset array called ds, for practical purposes, and define Tomato, Soil, and Fertilizer as categorical variables.
ds = fertilizer; ds.Tomato = nominal(ds.Tomato); ds.Soil = nominal(ds.Soil); ds.Fertilizer = nominal(ds.Fertilizer);
Fit a linear mixed-effects model, where Fertilizer and Tomato are the fixed-effects variables, and the mean yield varies by the block (soil type), and the plots within blocks (tomato types within soil types) independently.
lme = fitlme(ds,'Yield ~ Fertilizer * Tomato + (1|Soil) + (1|Soil:Tomato)');Predict the response values at the original design values. Display the first five predictions with the observed response values.
yhat = predict(lme); [yhat(1:5) ds.Yield(1:5)]
ans = 5×2
115.4788 104.0000
135.1455 136.0000
152.8121 158.0000
160.4788 174.0000
58.0839 57.0000
Plot Predictions vs. Observed Responses
Load the sample data.
load carsmallFit a linear mixed-effects model, with a fixed effect for Weight, and a random intercept grouped by Model_Year. First, store the data in a table.
tbl = table(MPG,Weight,Model_Year);
lme = fitlme(tbl,'MPG ~ Weight + (1|Model_Year)');Create predicted responses to the data.
yhat = predict(lme,tbl);
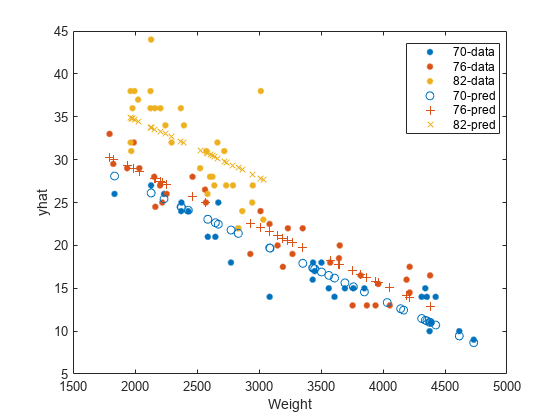
Plot the original responses and the predicted responses to see how they differ. Group them by model year.
figure() gscatter(Weight,MPG,Model_Year) hold on gscatter(Weight,yhat,Model_Year,[],'o+x') legend('70-data','76-data','82-data','70-pred','76-pred','82-pred') hold off
Predict Responses at Values in a New Dataset Array
Load the sample data.
load('fertilizer.mat');The dataset array includes data from a split-plot experiment, where soil is divided into three blocks based on the soil type: sandy, silty, and loamy. Each block is divided into five plots, where five different types of tomato plants (cherry, heirloom, grape, vine, and plum) are randomly assigned to these plots. The tomato plants in the plots are then divided into subplots, where each subplot is treated by one of four fertilizers. This is simulated data.
Store the data in a dataset array called ds, for practical purposes, and define Tomato, Soil, and Fertilizer as categorical variables.
ds = fertilizer; ds.Tomato = nominal(ds.Tomato); ds.Soil = nominal(ds.Soil); ds.Fertilizer = nominal(ds.Fertilizer);
Fit a linear mixed-effects model, where Fertilizer and Tomato are the fixed-effects variables, and the mean yield varies by the block (soil type), and the plots within blocks (tomato types within soil types) independently.
lme = fitlme(ds,'Yield ~ Fertilizer * Tomato + (1|Soil) + (1|Soil:Tomato)');Create a new dataset array with design values. The new dataset array must have the same variables as the original dataset array you use for fitting the model lme.
dsnew = dataset();
dsnew.Soil = nominal({'Sandy';'Silty'});
dsnew.Tomato = nominal({'Cherry';'Vine'});
dsnew. Fertilizer = nominal([2;2]);Predict the conditional and marginal responses at the original design points.
yhatC = predict(lme,dsnew);
yhatM = predict(lme,dsnew,'Conditional',false);
[yhatC yhatM]ans = 2×2
92.7505 111.6667
87.5891 82.6667
Predict Responses at the Values in New Design Matrices
Load the sample data.
load carbigFit a linear mixed-effects model for miles per gallon (MPG), with fixed effects for acceleration, horsepower, and cylinders, and potentially correlated random effects for intercept and acceleration grouped by model year.
First, prepare the design matrices for fitting the linear mixed-effects model.
X = [ones(406,1) Acceleration Horsepower]; Z = [ones(406,1) Acceleration]; Model_Year = nominal(Model_Year); G = Model_Year;
Now, fit the model using fitlmematrix with the defined design matrices and grouping variables.
lme = fitlmematrix(X,MPG,Z,G,'FixedEffectPredictors',.... {'Intercept','Acceleration','Horsepower'},'RandomEffectPredictors',... {{'Intercept','Acceleration'}},'RandomEffectGroups',{'Model_Year'});
Create the design matrices that contain the data at which to predict the response values. Xnew must have three columns as in X. The first column must be a column of 1s. And the values in the last two columns must correspond to Acceleration and Horsepower, respectively. The first column of Znew must be a column of 1s, and the second column must contain the same Acceleration values as in Xnew. The original grouping variable in G is the model year. So, Gnew must contain values for the model year. Note that Gnew must contain nominal values.
Xnew = [1,13.5,185; 1,17,205; 1,21.2,193];
Znew = [1,13.5; 1,17; 1,21.2]; % alternatively Znew = Xnew(:,1:2);
Gnew = nominal([73 77 82]);Predict the responses for the data in the new design matrices.
yhat = predict(lme,Xnew,Znew,Gnew)
yhat = 3×1
8.7063
5.4423
12.5384
Now, repeat the same for a linear mixed-effects model with uncorrelated random-effects terms for intercept and acceleration. First, change the original random effects design and the random effects grouping variables. Then, refit the model.
Z = {ones(406,1),Acceleration};
G = {Model_Year,Model_Year};
lme = fitlmematrix(X,MPG,Z,G,'FixedEffectPredictors',....
{'Intercept','Acceleration','Horsepower'},'RandomEffectPredictors',...
{{'Intercept'},{'Acceleration'}},'RandomEffectGroups',{'Model_Year','Model_Year'});Now, recreate the new random effects design, Znew, and the grouping variable design, Gnew, using which to predict the response values.
Znew = {[1;1;1],[13.5;17;21.2]};
MY = nominal([73 77 82]);
Gnew = {MY,MY};Predict the responses using the new design matrices.
yhat = predict(lme,Xnew,Znew,Gnew)
yhat = 3×1
8.6365
5.9199
12.1247
Compute Confidence Intervals for Predictions
Load the sample data.
load carbigFit a linear mixed-effects model for miles per gallon (MPG), with fixed effects for acceleration, horsepower, and cylinders, and potentially correlated random effects for intercept and acceleration grouped by model year. First, store the variables in a table.
tbl = table(MPG,Acceleration,Horsepower,Model_Year);
Now, fit the model using fitlme with the defined design matrices and grouping variables.
lme = fitlme(tbl,'MPG ~ Acceleration + Horsepower + (Acceleration|Model_Year)');Create the new data and store it in a new table.
tblnew = table(); tblnew.Acceleration = linspace(8,25)'; tblnew.Horsepower = linspace(min(Horsepower),max(Horsepower))'; tblnew.Model_Year = repmat(70,100,1);
linspace creates 100 equally distanced values between the lower and the upper input limits. Model_Year is fixed at 70. You can repeat this for any model year.
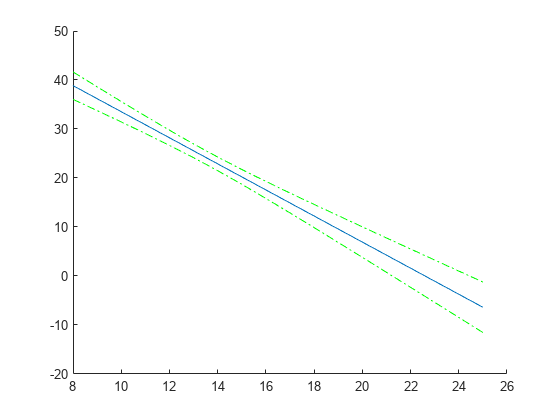
Compute and plot the predicted values and 95% confidence limits (nonsimultaneous).
[ypred,yCI,DF] = predict(lme,tblnew); figure(); h1 = line(tblnew.Acceleration,ypred); hold on; h2 = plot(tblnew.Acceleration,yCI,'g-.');
Display the degrees of freedom.
DF(1)
ans = 389
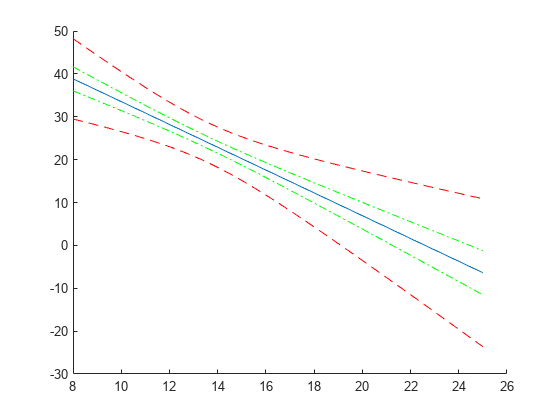
Compute and plot the simultaneous confidence bounds.
[ypred,yCI,DF] = predict(lme,tblnew,'Simultaneous',true); h3 = plot(tblnew.Acceleration,yCI,'r--');
Display the degrees of freedom.
DF
DF = 389
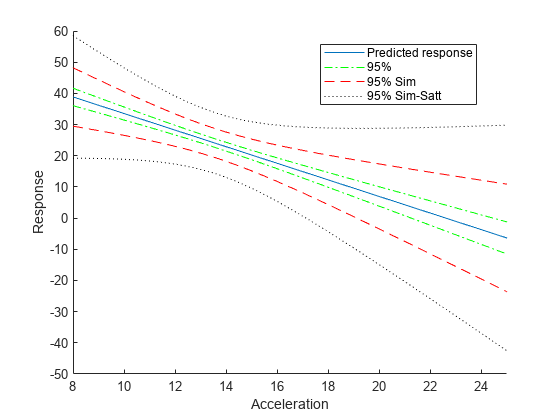
Compute the simultaneous confidence bounds using the Satterthwaite method to compute the degrees of freedom.
[ypred,yCI,DF] = predict(lme,tblnew,'Simultaneous',true,'DFMethod','satterthwaite'); h4 = plot(tblnew.Acceleration,yCI,'k:'); hold off xlabel('Acceleration') ylabel('Response') ylim([-50,60]) xlim([8,25]) legend([h1,h2(1),h3(1),h4(1)],'Predicted response','95%','95% Sim',... '95% Sim-Satt','Location','Best')
Display the degrees of freedom.
DF
DF = 3.6001
Input Arguments
lme — Linear mixed-effects model
LinearMixedModel object
Linear mixed-effects model, specified as a LinearMixedModel object constructed using fitlme or fitlmematrix.
tblnew — New input data
table | dataset array
New input data, which includes the response variable, predictor
variables, and grouping
variables, specified as a table or dataset array. The predictor
variables can be continuous or grouping variables. tblnew must
have the same variables as in the original table or dataset array
used to fit the linear mixed-effects model lme.
Xnew — New fixed-effects design matrix
n-by-p matrix
New fixed-effects design matrix, specified as an n-by-p matrix,
where n is the number of observations and p is
the number of fixed predictor variables. Each row of X corresponds
to one observation and each column of X corresponds
to one variable.
Data Types: single | double
Znew — New random-effects design
n-by-q matrix | cell array of length R
New random-effects design, specified as an n-by-q matrix
or a cell array of R design matrices Z{r},
where r = 1, 2, ..., R. If Znew is
a cell array, then each Z{r} is an n-by-q(r)
matrix, where n is the number of observations,
and q(r) is the number of random
predictor variables.
Data Types: single | double | cell
Gnew — New grouping variable or variables
vector | cell array of grouping variables of length R
New grouping variable or variables, specified
as a vector or a cell array, of length R, of grouping
variables with the same levels or groups as the original grouping
variables used to fit the linear mixed-effects model lme.
Data Types: single | double | categorical | logical | char | string | cell
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: ypred = predict(lme,'Conditional',false);
Alpha — Significance level
0.05 (default) | scalar value in the range 0 to 1
Significance level, specified as the comma-separated pair consisting of
'Alpha' and a scalar value in the range 0 to 1. For a value α,
the confidence level is 100*(1–α)%.
For example, for 99% confidence intervals, you can specify the confidence level as follows.
Example: 'Alpha',0.01
Data Types: single | double
Conditional — Indicator for conditional predictions
true (default) | false
Indicator for conditional predictions,
specified as the comma-separated pair consisting of 'Conditional' and
one of the following.
true | Contributions from both fixed effects and random effects (conditional) |
false | Contribution from only fixed effects (marginal) |
Example: 'Conditional,false
DFMethod — Method for computing approximate degrees of freedom
'residual' (default) | 'satterthwaite' | 'none'
Method for computing approximate degrees of freedom to use in
the confidence interval computation, specified as the comma-separated
pair consisting of 'DFMethod' and one of the following.
'residual' | Default. The degrees of freedom are assumed to be constant and equal to n – p, where n is the number of observations and p is the number of fixed effects. |
'satterthwaite' | Satterthwaite approximation. |
'none' | All degrees of freedom are set to infinity. |
For example, you can specify the Satterthwaite approximation as follows.
Example: 'DFMethod','satterthwaite'
Simultaneous — Type of confidence bounds
false (default) | true
Type of confidence bounds, specified as the comma-separated
pair consisting of 'Simultaneous' and one of the
following.
false | Default. Nonsimultaneous bounds. |
true | Simultaneous bounds. |
Example: 'Simultaneous',true
Prediction — Type of prediction
'curve' (default) | 'observation'
Type of prediction, specified as the comma-separated pair consisting
of 'Prediction' and one of the following.
'curve' | Default. Confidence bounds for the predictions based on the fitted function. |
'observation' | Variability due to observation error for the new observations is also included in the confidence bound calculations and this results in wider bounds. |
Example: 'Prediction','observation'
Output Arguments
ypred — Predicted responses
vector
Predicted responses, returned as a vector. ypred can
contain the conditional or marginal responses, depending on the value
choice of the 'Conditional' name-value pair argument.
Conditional predictions include contributions from both fixed and
random effects.
ypredCI — Point-wise confidence intervals
two-column matrix
Point-wise confidence intervals for the predicted values, returned
as a two-column matrix. The first column of yCI contains
the lower bounds, and the second column contains the upper bound.
By default, yCI contains the 95% confidence intervals
for the predictions. You can change the confidence level using the Alpha name-value
pair argument, make them simultaneous using the Simultaneous name-value
pair argument, and also make them for a new observation rather than
for the curve using the Prediction name-value
pair argument.
DF — Degrees of freedom
vector | scalar value
Degrees of freedom used in computing the confidence intervals, returned as a vector or a scalar value.
If the
'Simultaneous'name-value pair argument isfalse, thenDFis a vector.If the
'Simultaneous'name-value pair argument istrue, thenDFis a scalar value.
More About
Conditional and Marginal Predictions
A conditional prediction includes contributions from both fixed and random effects, whereas a marginal model includes contribution from only fixed effects.
Suppose the linear mixed-effects model lme has
an n-by-p fixed-effects design
matrix X and an n-by-q random-effects
design matrix Z. Also, suppose the estimated p-by-1
fixed-effects vector is ,
and the q-by-1 estimated best linear unbiased predictor
(BLUP) vector of random effects is .
The predicted conditional response is
which corresponds to the 'Conditional','true' name-value
pair argument.
The predicted marginal response is
which corresponds to the 'Conditional','false' name-value
pair argument.
When making predictions, if a particular grouping variable has
new levels (1s that were not in the original data), then the random
effects for the grouping variable do not contribute to the 'Conditional' prediction
at observations where the grouping variable has new levels.
Version History
Introduced in R2013b
See Also
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.

Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list:
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)