1-D Adaptive Thresholding of Wavelet Coefficients
This section takes you through the features of local thresholding of wavelet coefficients for 1-D signals or data. This capability is available through Wavelet Analyzer app:
Wavelet Denoising 1-D
Wavelet Compression 1-D
SWT Denoising 1-D
Regression Estimation 1-D
Density Estimation 1-D
This tool allows you to define, level by level, time-dependent (x-axis-dependent) thresholds, and then increase the capability of the denoising strategies handling nonstationary variance noise. More precisely, the model assumes that the observation is equal to the interesting signal superimposed on noise. The noise variance can vary with time. There are several different variance values on several time intervals. The values as well as the intervals are unknown. This section will use one of Wavelet Analyzer app tools (SWT Denoising 1-D) to illustrate this capability. The behavior of all the above-mentioned tools is similar.
1-D Local Thresholding Using the Wavelet Analyzer App
From the MATLAB® prompt, type
waveletAnalyzer.The Wavelet Analyzer appears.
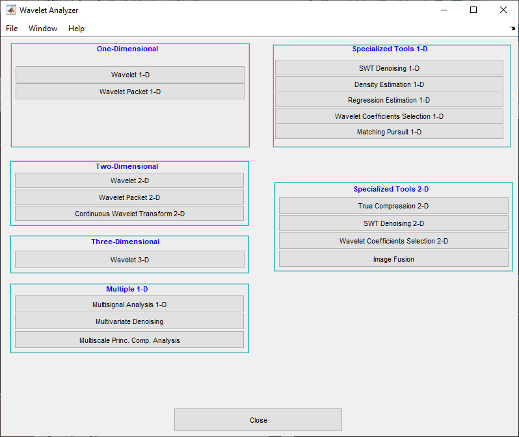
Click the SWT Denoising 1-D menu item.
The discrete stationary wavelet transform denoising tool for 1-D signals appears.
Load data.
At the MATLAB command prompt, type
In the SWT Denoising 1-D tool, select File > Import from Workspace. When the Import from Workspace dialog box appears, select theload nblocr1;
nblocr1variable. Click OK to import the noisy blocks signal with two change points in the noise variance located at positions 200 and 600.Perform signal decomposition.
Select the
db1wavelet from the Wavelet menu and select 5 from the Level menu, and then click the Decompose Signal button. After a pause for computation, the tool displays the stationary wavelet approximation and detail coefficients of the decomposition.Accept the defaults of Fixed form soft thresholding and Unscaled white noise. Click the Denoise button.
The result is quite satisfactory, but seems to be oversmoothed when the signal is irregular.
Select hard for the thresholding mode instead of soft, and then click the Denoise button.
The result is not satisfactory. The denoised signal remains noisy before position 200 and after position 700. This illustrates the limits of the classical denoising strategies. In addition, the residuals obtained during the last trials clearly suggest to try a local thresholding strategy.
Generate interval-dependent thresholds.
Click the Int. dependent threshold Settings button located at the bottom of the thresholding method frame. A new window titled Int. Dependent Threshold Settings for figure ... appears.
Click the Generate button. After a pause for computation, the tool displays the default intervals associated with adapted thresholds.
Three intervals are proposed. Since the variances for the three intervals are very different, the optimization program easily detects the correct structure. Nevertheless, you can visualize the intervals proposed for a number of intervals from 1 to 6 using the Select Number of Intervals menu (which replaces the Generate button). Using the default intervals automatically propagates the interval delimiters and associated thresholds to all levels.
Denoise with Interval-Dependent Thresholds
Click the Close button in the Int. Dependent Threshold Settings for ... window. When the Update thresholds dialog box appears, click Yes. The SWT Denoising 1-D main window is updated. The sliders located to the right of the window control the level and interval dependent thresholds. For a given interval, the threshold is indicated by yellow dotted lines running horizontally through the graphs on the left of the window. The red dotted lines running vertically through the graphs indicate the interval delimiters. Next click the Denoise button.
Modifying Interval Dependent Thresholds
The thresholds can be increased to keep only the highest values of the wavelet coefficients at each level. Do this by dragging the yellow lines directly on the graphs on the left of the window, or using the View Axes button (located at the bottom of the screen near the Close button), which allows you to see each axis in full size. Another way is to edit the thresholds by selecting the interval number located near the sliders and typing the desired value.
Note that you can also change the interval limits by holding down the left mouse button over the vertical dotted red lines, and dragging them.
You can also define your own interval dependent strategy. Click the Int. dependent threshold settings button. The Int. Dependent Threshold Settings for ... window appears again. We shall explore this window for a little while. Click the Delete button, so that the interval delimiters disappear. Double click the left mouse button to define new interval delimiters; for example at positions 300 and 500 and adjust the thresholds manually. Each level must be considered separately using the Level menu for adjusting the thresholds. The current interval delimiters can be propagated to all levels by clicking the Propagate button. So click the Propagate button. Adjust the thresholds for each level, one by one. At the end, click the Close button of the Int. Dependent Threshold settings for ... window. When the Update thresholds dialog box appears, click Yes. Then click the denoise button.
Note that
By double-clicking again on an interval delimiter with the left mouse button, you delete it.
You can move the interval delimiters (vertical red dotted lines) and the threshold levels (horizontal yellow dotted lines) by holding down the left mouse button over these lines and dragging them.
The maximum number of interval delimiters at each level is 10.
Examples of Denoising with Interval Dependent Thresholds.
From the File menu, choose the Example Analysis > Noisy Signals - Interval Dependent Noise
Variance > option. From the drop down men, choose
with haar at level 4 ---> Elec. consumption — 3
intervals. The proposed items contain, in addition to the
usual information, the “true” number of intervals. You can then
experiment with various signals for which local thresholding is needed.
Importing and Exporting Information from the Wavelet Analyzer App
The tool lets you save the denoised signal to disk. The toolbox creates a MAT-file in the current folder with a name you choose.
To save the denoised signal from the present denoising process, use the menu
option File > Save denoised Signal. A dialog box
appears that lets you specify a folder and filename for storing the signal. Type the
name dnelec. After saving the signal data to the file
dnelec.mat, load the variables into your workspace:
load dnelec whos
| Name | Size | Bytes | Class |
|---|---|---|---|
dnelec | 1x2000 | 16000 | double array |
thrParams | 1x4 | 656 | cell array |
wname | 1x4 | 8 | char array |
The denoised signal is given by dnelec. In addition, the
parameters of the denoising process are given by the wavelet name contained in
wname:
wname
wname =
haar
and the level dependent thresholds contained in thrParams,
which is a cell array of length 4 (the level of the decomposition). For
i from 1 to 4, thrParams{i} is an array
nbintx3 (where nbint is the number of
intervals, here 3), and each row contains the lower and upper bounds of the interval
of thresholding and the threshold value. For example, for level 1,
thrParams{1}
ans =
1.0e+03 *
0.0010 0.0980 0.0060
0.0980 1.1240 0.0204
1.1240 2.0000 0.0049
