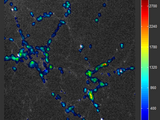
HotPlot, view a 2D variable as hue over 2D data
I needed a way to visualize a subset of a 2D variable derived from a 2D dataset, _together_ with the original data and in its original coordinates. Since a plain 3D plot decouples all this info (and becomes illegible for data containing very noisy regions) I decided to show the calculated 2D descriptor as hue variation on the original 2D data (represented as gray level).
It is a suggestive way of visualizing data and especially effective when the 2D subset is a lot smaller than the original 2D data (for instance when the variable to plot is uninteresting for the most part and it would only skew the z-scaling).
Following the discussions on the FileExchange and Newsgroup it seemed to me that more people might benefit from this, so I decided to post it.
execute:
>> imgRGB = hotplot(BACKGND, FOREGND, POSITION, 1);
when FOREGND is an N-length list of values and POSITION is an N-by-2 list of coordinates (in the BACKGND referential) ;
or execute:
>> imgRGB = hotplot(BACKGND, FOREGND, MASK, 1);
when BACKGND and FOREGND are both [M x N] and MASK is a boolean, obtained for example as
>> MASK = FOREGND > SomeThreshold;
or run testHotPlot.m to visualise a sample data set; the physical meaning of this dataset is :
FOREGND - neural impulse energy
BACKGND - average neural potential
for ideas, suggestions, etc. mail me at
tudima at y a h o o dot com
Cite As
tudor dima (2024). HotPlot, view a 2D variable as hue over 2D data (https://www.mathworks.com/matlabcentral/fileexchange/25301-hotplot-view-a-2d-variable-as-hue-over-2d-data), MATLAB Central File Exchange. Retrieved .
MATLAB Release Compatibility
Platform Compatibility
Windows macOS LinuxCategories
- MATLAB > Graphics > 2-D and 3-D Plots > Line Plots >
Tags
Community Treasure Hunt
Find the treasures in MATLAB Central and discover how the community can help you!
Start Hunting!Discover Live Editor
Create scripts with code, output, and formatted text in a single executable document.