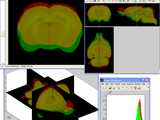
GUI for exploration of 3D images (stacks)
You are now following this Submission
- You will see updates in your followed content feed
- You may receive emails, depending on your communication preferences
A graphical user interface for display and exploration of 3D images of various types. Manages grayscale, RGB, label images. Allows orthoslices display, isosurface reconstruction, changing intensity range or look-up table, and basic type conversions.
Part of the "MatImage" toolbox (http://github.com/dlegland/matImage )
Requires the GUI Layout Toolbox to be installed
Cite As
David Legland (2026). Slicer (https://www.mathworks.com/matlabcentral/fileexchange/27983-slicer), MATLAB Central File Exchange. Retrieved .
Acknowledgements
Inspired by: tiffread2.m, GUI Layout Toolbox, myslicer - make mouse-interactive slices of a 3-D volume
Inspired: pcolor3
General Information
- Version 1.0.3.0 (190 KB)
MATLAB Release Compatibility
- Compatible with any release
Platform Compatibility
- Windows
- macOS
- Linux
| Version | Published | Release Notes | Action |
|---|---|---|---|
| 1.10.0.0 | update to work with GuiLayout Toolbox 2, and package as a Matlab Application |
||
| 1.9.0.0 | better support of the different image types (binary, label...), more options for displaying orthoslices and histograms, added demo images, read offset from mhd files |
||
| 1.7.0.0 | Updated to include an App file for R2012b |
||
| 1.6.0.0 | Switched the interface to GUILayout Toolbox. Lot of code rewritting... |
||
| 1.5.0.0 | better control on size/spacing when importing metaImage format |
||
| 1.4.0.0 | fix bug when opening without arguments |
||
| 1.3.0.0 | added display of orthoslices, enhanced loading of new images, enhance display of image infos (name, pixel value...) |
||
| 1.1.0.0 | added options to choose grayscale extent, lut management, and fixed several bugs. |
||
| 1.0.3.0 | better management of older releases, add support of isosurface computation and for z-profile display |
||
| 1.0.2.0 | fix some GUI bugs. |
||
| 1.0.1.0 | Better management of GUI Layout v2.1 |
||
| 1.0.0.0 | fix bug when opening with empty image |