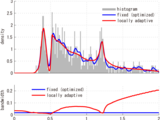
ssvkernel(x,tin)
First thing to do:
Run a tutorial code, tutorial.m
Documentation:
http://2000.jukuin.keio.ac.jp/shimazaki/res/kernel.html
see also sskernel for optimization of a fixed kernel bandwidth and sshist for histogram optimization.
% [y,t,optw,gs,C,confb95,yb] = ssvkernel(x,t,W)
%
% Function `ssvkernel' returns an optimized kernel density estimate
% using a Gauss kernel function with bandwidths locally adapted to the data.
%
% Examples:
% >> x = 0.5-0.5*log(rand(1,1e3)); t = linspace(0,3,500);
% >> [y,t,optw] = ssvkernel(x,t);
% This example produces a vector of kernel density estimates, y, at points
% specified in a vector t, using locally adaptive bandwidths, optw
% (a standard deviation of a normal density function).
%
% >> ssvkernel(x);
% By calling the function without output arguments, the estimated density
% is displayed.
%
% Input arguments:
% x: Sample data vector.
% tin (optinal):
% Points at which estimation are computed.
% W (optinal):
% A vector of kernel bandwidths.
% If W is provided, the optimal bandwidth is selected from the
% elements of W.
% * Do not search bandwidths smaller than a sampling resolution of data.
% If W is not provided, the program searches the optimal bandwidth
% using a golden section search method.
%
% Output arguments:
% y: Estimated density
% t: Points at which estimation was computed.
% The same as tin if tin is provided.
% (If the sampling resolution of tin is smaller than the sampling
% resolution of the data, x, the estimation was done at smaller
% number of points than t. The results, t and y, are obtained by
% interpolating the low resolution sampling points.)
% optw: Optimal kernel bandwidth.
% gs: Stiffness constants of the variable bandwidth examined.
% The stifness constant is defined as a ratio of the optimal fixed
% bandwidth to a length of a local interval in which a fixed-kernel
% bandwidth optimization was performed.
% C: Cost functions of stiffness constants.
% conf95:
% Bootstrap confidence intervals.
% yb: Booststrap samples.
%
%
% Usage:
% >> [y,t,optw] = ssvkernel(x);
% When t is not given in the input arguments, i.e., the output argument t
% is generated automatically.
%
% >> W = linspace(0.01,1,20);
% >> [y,t,optw] = ssvkernel(x,t,W);
% The optimal bandwidth is selected from the elements of W.
%
% >> [y,t,optw,confb95,yb] = ssvkernel(x);
% This additionally computes 95% bootstrap confidence intervals, confb95.
% The bootstrap samples are provided as yb.
%
%
% Optimization principle:
% The optimization is based on a principle of minimizing
% expected L2 loss function between the kernel estimate and an unknown
% underlying density function. An assumption is merely that samples
% are drawn from the density independently each other.
%
% The locally adaptive bandwidth is obtained by iteratively computing
% optimal fixed-size bandwidths wihtihn local intervals. The optimal
% bandwidths are selected such that they are selected in the intervals
% that are \gamma times larger than the optimal bandwidths themselves.
% The paramter \gamma was optimized by minimizing the L2 risk estimate.
%
% The method is described in
% Hideaki Shimazaki and Shigeru Shinomoto
% Kernel Bandwidth Optimization in Spike Rate Estimation
% Journal of Computational Neuroscience 2010
% http://dx.doi.org/10.1007/s10827-009-0180-4
%
% * Instead of a Gaussian window described in the paper, a boxcar window is
% used in this program.
%
% For more information, please visit
% http://2000.jukuin.keio.ac.jp/shimazaki/res/kernel.html
%
% See also SSKERNEL, SSHIST
%
%
% Hideaki Shimazaki
% http://2000.jukuin.keio.ac.jp/shimazaki
Cite As
Hideaki Shimazaki (2024). ssvkernel(x,tin) (https://www.mathworks.com/matlabcentral/fileexchange/37374-ssvkernel-x-tin), MATLAB Central File Exchange. Retrieved .
MATLAB Release Compatibility
Platform Compatibility
Windows macOS LinuxCategories
- Sciences > Neuroscience > Frequently-used Algorithms >
- Sciences > Neuroscience > Cellular Neuroscience > Electrophysiology >
Tags
Community Treasure Hunt
Find the treasures in MATLAB Central and discover how the community can help you!
Start Hunting!Discover Live Editor
Create scripts with code, output, and formatted text in a single executable document.
| Version | Published | Release Notes | |
|---|---|---|---|
| 1.3.0.0 | Bug fix: fixed a problem for large values
|
||
| 1.2.0.0 | fixed issues on compatibility to older versions of matlab. performance optimization. |
||
| 1.1.0.0 | correcting url in Description |
||
| 1.0.0.0 |