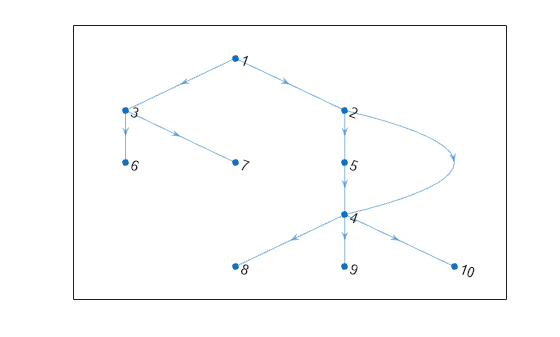
isdag
Determine if graph is acyclic
Syntax
Description
tf = isdag( returns logical
G)1 (true) if G is a
directed acyclic graph;
otherwise, it returns logical 0
(false).
Examples
Input Arguments
More About
Version History
Introduced in R2015b
See Also
toposort | reordernodes | digraph | hascycles