proteinplot
Open Protein Plot window to investigate properties of amino acid sequence
Description
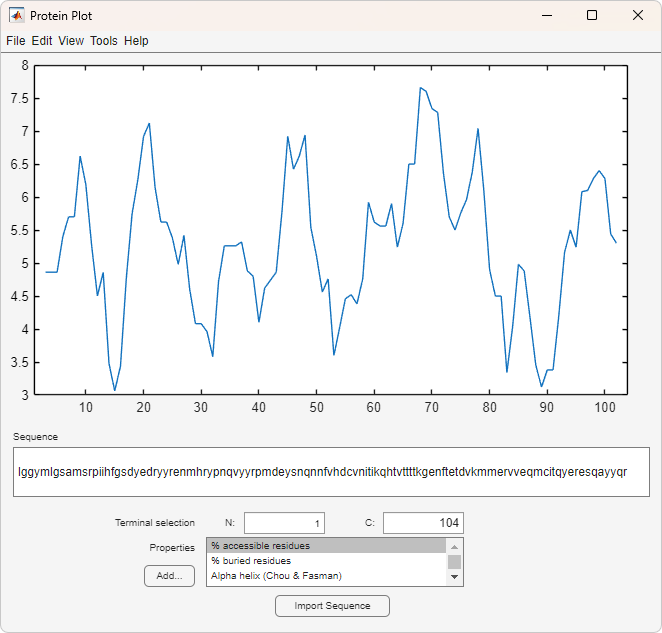
proteinplot opens the Protein Plot window that enables you to
analyze and compare properties of a single amino acid sequence. It displays smoothed line
plots of various properties such as the hydrophobicity of the amino acids in the
sequence.
proteinplot( opens the Protein Plot
window and loads SeqAA)SeqAA, an amino acid sequence, into the window.
You can analyze and compare properties of an amino acid sequence from the MATLAB® command line also by using the proteinpropplot function.
Examples
Input Arguments
Version History
Introduced before R2006a
See Also
aacount | atomiccomp | molweight | pdbdistplot | proteinpropplot | yyaxis