Detect Anomalies in Pills During Live Image Acquisition
This example shows how to detect anomalies in pills during live image acquisition. In this example, you modify the pretrained neural network from Detect Image Anomalies Using Explainable FCDD Network (Computer Vision Toolbox).
You first create a data set of images of normal pills and pills with anomalies, using image acquisition hardware. Then, you use this data set to calibrate and test the pretrained neural network.
Next, you capture live images of pills and use the trained anomaly detector to classify the pill as normal or anomalous. The live image preview also displays a text label that indicates the probability that each pixel is normal or anomalous.
Finally, you deploy the trained anomaly detection model and live image acquisition using MATLAB® Compiler™. Once the script is deployed as a standalone executable, you can launch it on any machine without a MATLAB license.
This example uses a webcam to capture the pill images for calibration, testing, and live pill image classification. However, you can run this example using any image acquisition hardware that is supported by the Image Acquisition Toolbox™.
In addition to the required MATLAB toolboxes, you must also have the following add-ons installed to run this example.
Image Acquisition Toolbox Support Package for OS Generic Video Interface
Computer Vision Toolbox™ Automated Visual Inspection Library
Download Pretrained Network
This example uses a pretrained network that is trained to detect pill anomalies. Download this network.
trainedPillAnomalyDetectorNet_url = ... "https://ssd.mathworks.com/supportfiles/vision/data/trainedFCDDPillAnomalyDetectorSpkg.zip"; downloadTrainedNetwork(trainedPillAnomalyDetectorNet_url,pwd); net = load(fullfile(pwd,"folderForSupportFilesInceptionModel","trainedPillFCDDNet.mat")); detector = net.detector;
Prepare Data Set for Calibration and Testing
You use a pretrained pill anomaly detector because the images used to train it are similar to the images captured and classified in this example. Rather than retrain the existing model from scratch, you capture images for calibration and testing. If your workflow requires a visual inspection model for a different application, refer to Detect Image Anomalies Using Explainable FCDD Network (Computer Vision Toolbox) for more information on how to train the model from scratch.
Set Up Image Acquisition Hardware
Connect to your camera using the videoinput function and specify its properties. This example uses the OS generic video interface to acquire images from a Logitech® webcam with the YUV2_352x288 video format.
vid = videoinput("winvideo","1","YUY2_352x288"); vid.ReturnedColorSpace = 'rgb'; vid.FramesPerTrigger = 70;
Get the source properties of the videoinput object. Specify device-specific properties to adjust for optimal image data collection. The values you specify for these properties depend on the camera hardware and the surrounding environment that images are being captured in.
src = getselectedsource(vid);
Acquire Data
After setting up and configuring your image acquisition hardware, you can collect images. Collect data in immediate acquisition mode. For more information about trigger types, see Specifying the Trigger Type.
start(vid); wait(vid);
After you finish capturing images, save the acquired image frames to MATLAB workspace and save the frames as TIF files using imwrite.
img = getdata(vid,70); for f=1:vid.FramesAcquired imwrite(img(:,:,:,f), "image"+f+".tif"); end delete(vid);
In the current working directory create a folder titled ImageData. Within this folder create two new folders titled normal and dirt, then move images of normal and anomalous pills, respectively, into these folders.
The original data set used in this example contained 40 normal images and 30 dirt images.
These two images show an example from each class. A normal pill with no defects is on the left and a pill contaminated with dirt is on the right. The properties of these images align closely with the properties of the training data set from the Detect Image Anomalies Using Explainable FCDD Network (Computer Vision Toolbox) example.
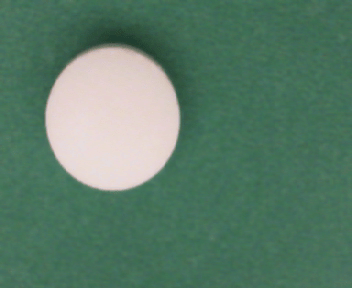


Load and Preprocess Data
Create an imageDataStore object, imds, to manage the collection of image files from both normal and dirt image classes.
imageDir = fullfile(pwd,"ImageData"); imds = imageDatastore(imageDir,IncludeSubfolders=true,LabelSource="foldernames");
Split imds into calibration and testing sets (imdsCal and imdsTest, respectively). The images for calibration are used to determine a threshold value for the classifier. The classifier labels images with anomaly scores above the threshold as anomalous. The remaining images are for testing and are used to evaluate the classifier.
normalTrainRatio = 0; anomalyTrainRatio = 0; normalCalRatio = 0.70; anomalyCalRatio = 0.60; normalTestRatio = 1 - (normalTrainRatio + normalCalRatio); anomalyTestRatio = 1 - (anomalyTrainRatio + anomalyCalRatio); rng(0); [imdsTrain,imdsCal,imdsTest] = splitAnomalyData(imds,["dirt"],... NormalLabelsRatio=[normalTrainRatio normalCalRatio normalTestRatio],... AnomalyLabelsRatio=[anomalyTrainRatio anomalyCalRatio anomalyTestRatio]);
Splitting anomaly dataset
-------------------------
* Finalizing... Done.
* Number of files and proportions per class in all the datasets:
Input Train Validation Test
NumFiles Ratio NumFiles Ratio NumFiles Ratio NumFiles Ratio
_____________________________ _________________ _____________________________ _________________
dirt 30 0.428571428571429 0 0 18 0.391304347826087 12 0.5
normal 40 0.571428571428571 0 0 28 0.608695652173913 12 0.5
Add binary labels to normal and dirt images in both calibration and test data sets by using the transform function with the operations specified by the addLabelData helper function.
dsCal = transform(imdsCal,@addLabelData,IncludeInfo=true); dsTest = transform(imdsTest,@addLabelData,IncludeInfo=true);
Visualize a sample of calibration data that contains both normal and anomalous images.
numObs = length(imdsCal.Labels); idx = randperm(numObs,9); montage(imdsCal,Indices=idx)
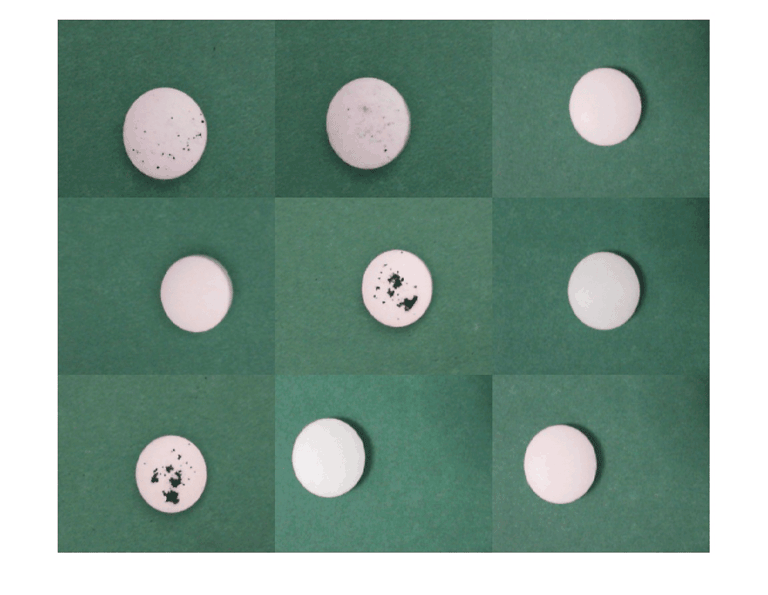

Calibrate and Test Classification Model
Select Anomaly Threshold
Use the calibration data set to select an anomaly score threshold for the anomaly detector. The detector classifies images based on whether their scores are above or below the threshold value. You use both normal and anomalous images to determine the threshold.
Get the mean anomaly score and ground truth label for each image in the calibration set and plot a histogram of scores.
scores = predict(detector,dsCal); labels = imdsCal.Labels ~= "normal"; numBins = 20; [~,edges] = histcounts(scores,numBins); figure hold on hNormal = histogram(scores(labels==0),edges); hAnomaly = histogram(scores(labels==1),edges); hold off legend([hNormal,hAnomaly],"Normal","Anomaly") xlabel("Mean Anomaly Score") ylabel("Counts")
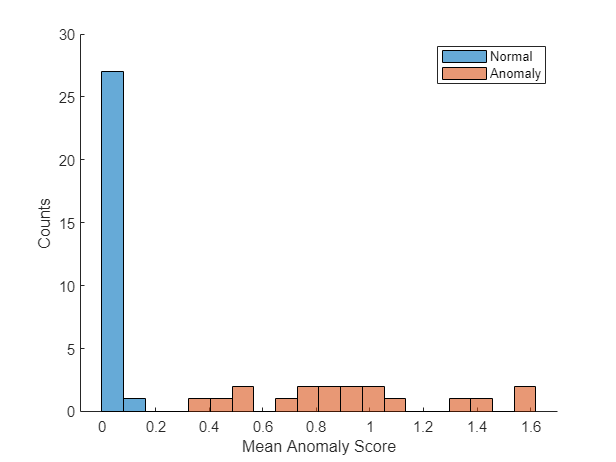
Calculate the optimal anomaly threshold using the anomalyThreshold (Computer Vision Toolbox) function. Specify the first two input arguments as the ground truth labels, labels, and predicted anomaly scores, scores, for the calibration data set. Specify the third input argument as true because true positive anomalous images have a labels value of true.
thresh = anomalyThreshold(labels,scores,true);
Set the Threshold property of the anomaly detector to the optimal value.
detector.Threshold = thresh;
Evaluate Classification Model
Classify each image in the test set as either normal or anomalous using the anomaly detector, detector.
testSetOutputLabels = classify(detector,dsTest);
Get the ground truth labels of each test image.
testSetTargetLabels = dsTest.UnderlyingDatastores{1}.Labels;Evaluate the anomaly detector by calculating performance metrics by using the evaluateAnomalyDetection (Computer Vision Toolbox) function. The function calculates several metrics that evaluate the accuracy, precision, sensitivity, and specificity of the detector for the test data set.
metrics = evaluateAnomalyDetection(testSetOutputLabels,testSetTargetLabels,"dirt"); Evaluating anomaly detection results
------------------------------------
* Finalizing... Done.
* Data set metrics:
GlobalAccuracy MeanAccuracy Precision Recall Specificity F1Score FalsePositiveRate FalseNegativeRate
______________ ____________ _________ ______ ___________ _______ _________________ _________________
1 1 1 1 1 1 0 0
The ConfusionMatrix property of metrics contains the confusion matrix for the test set. Extract the confusion matrix and display a confusion plot. The recalibrated classification model with the anomaly threshold selected produces very few false positives and false negatives.
M = metrics.ConfusionMatrix{:,:};
confusionchart(M,["Normal","Anomaly"])
acc = sum(diag(M)) / sum(M,"all");
title("Accuracy: "+acc)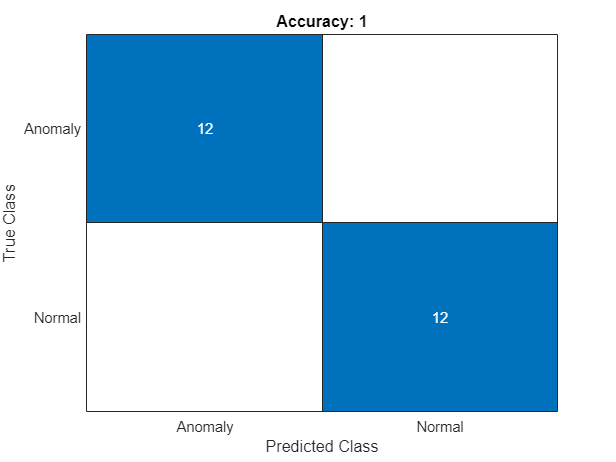
For more information about explainable classification decisions, see the Detect Image Anomalies Using Explainable FCDD Network (Computer Vision Toolbox) example.
Save the detector to a MAT file for later use in MATLAB or in deployment workflows.
save('livePillDetector.mat','detector')
Classify Live Images of Pills
You can use a custom video preview from a camera to display live classification of pills as normal or anomalous. This section demonstrates how to create a custom preview update callback function to process live images, identify individual pills, classify each pill, and show classification results overlaid with the camera preview video. You can find the code from this section as well as the update preview callback function attached to this example as supporting files.
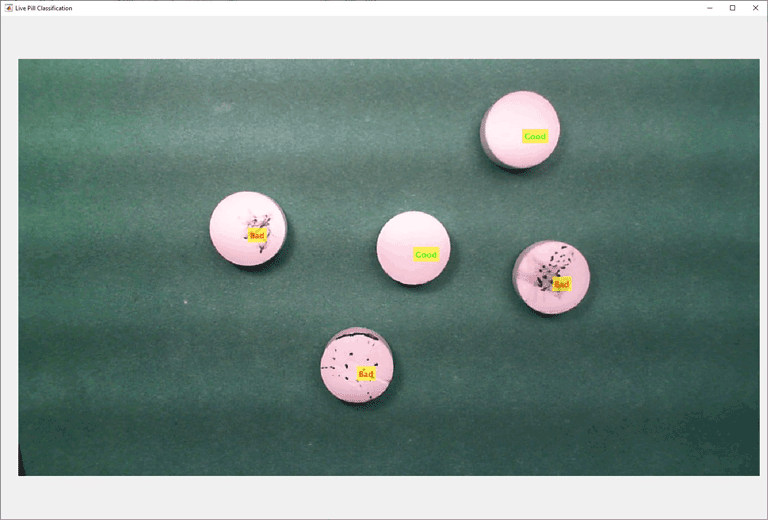
Acquire images using a webcam with the videoinput interface at a resolution of 1280-by-720. Then, use a custom preview update callback function to identify each pill using the imfindcircles function, since the pills are circular. Crop the section of the camera image around each pill and pass it to the detector for classification. Based on the classification decision, display a label on each pill indicating whether it is a normal or anomalous pill. Normal pills are labeled as Good and anomalous pills are labeled as Bad in the preview window.
Load the anomaly detector.
load('livePillDetector.mat');Create a connection to the webcam and specify appropriate values for the device-specific properties. The following device-specific property values are optimized for this hardware setup used originally for this example. Your setup might require a different configuration.
vid = videoinput('winvideo',1,'YUY2_1280x720'); vid.ReturnedColorSpace = "rgb"; nBands = vid.NumberOfBands; vidRes = vid.ROIPosition; imWidth = vidRes(3); imHeight = vidRes(4); src = getselectedsource(vid); src.ExposureMode = "manual"; src.Exposure = -11; src.FocusMode = "manual"; src.Focus = 11; src.WhiteBalanceMode = "manual"; src.WhiteBalance = 2800; src.FrameRate = "20.0000";
Create a figure window and axes in the figure. Then create the image object in which you want to display the video preview data. The size of the image object must match the dimensions of the video frames. Add a scroll panel to the image.
hFig = uifigure('Toolbar','none',... 'Menubar', 'none',... 'NumberTitle','Off',... 'AutoResizeChildren','Off',... 'Name','Live Pill Classification'); hAxis = uiaxes(hFig); disableDefaultInteractivity(hAxis); hImage = image(zeros(imHeight, imWidth, nBands),'Parent',hAxis); imscrollpanel(hFig,hImage);
Configure the update preview window callback function and the detector as application-defined data on the image object. To define the mypreview_fcn callback function, refer to the Perform Custom Processing of Preview Data section.
setappdata(hImage,'UpdatePreviewWindowFcn',@mypreview_fcn); setappdata(hImage,'Detector',detector);
Preview the video data in the custom figure window.
preview(vid,hImage);

Perform Custom Processing of Preview Data
Create a callback function mypreview_fcn that processes each frame and identifies whether pills are normal or dirt. This callback function is called for each acquired frame when you call the preview function.
The mypreview_fcn callback function performs the following:
Processes each frame that is captured and determines the number of circles in it using
imfindcircles. Since the pills are circular, theimfindcirclesfunction identifies the number of pills based on the specified radius range. Determine the radius range by measuring the radius of a pill in pixels using the Distance tool on a captured image. For more information on how to use the Distance tool, see Measure Distances and Areas Using Image Viewer App.Crops the image around each detected circle and passes the cropped image to the classifier for classification.
Adds a text label to the center of each circle that identifies the pill as
GoodorBadbased on the classification decision.Repeats the cropping and labeling steps for every circle detected.
Updates the image in the custom preview window.
function mypreview_fcn(obj,event,hImage) persistent detector; persistent ImgSize; if isempty(detector) && isempty(ImgSize) % Anomaly detector detector = getappdata(hImage,'Detector'); % Width and height of images used during calibration ImgSize = [352 288]; end % Find circles (pills) in the current image frame img = event.Data; radiusRange = [30 80]; circles = imfindcircles(img,radiusRange); % Use classifier if circles are found if ~isempty(circles) % Loop through all circles for i = 1:size(circles,1) % Crop image around the circle (pill) % [pos1, pos2] - top-left corner pixel position % ImgSize - width and height from [pos1, pos2] pos1 = max(circles(i,1)-ImgSize(1)/2,0); pos2 = max(circles(i,2)-ImgSize(2)/2,0); croppedImage = imcrop(img,[pos1, pos2,ImgSize(1),ImgSize(2)]); if ~isempty(croppedImage) % Classify cropped image and assign a text label in the % center of the circle decision = classify(detector, croppedImage); if decision img = insertText(img,circles(i,:),"Bad", TextColor="Red",FontSize=14); else img = insertText(img,circles(i,:),"Good", TextColor="Green",FontSize=14); end end end end hImage.CData = img; end
This code is expected to classify images correctly if the camera's field of view contains one pill. If the field of view contains multiple pills, the distance between the pills must be large enough so that the cropped images do not contain more than one pill.
Deploy Live Image Classification as Standalone Application
You can deploy the live image classification of pills as a standalone application using the Standalone Application Compiler (MATLAB Compiler) app. You must have a license for MATLAB Compiler. Once you create the standalone application, you can deploy it to multiple machines without additional MATLAB licenses.
You must do the following in the Application Compiler app to create the standalone application.
Add
customGUIForLiveClassification.mas the main file for the project.Add the Image Acquisition Toolbox™ Support Package for OS Generic Video Interface and Computer Vision Toolbox™ Automated Visual Inspection Library from the suggested list of support packages.
See Also
videoinput | getdata | imwrite | imageDatastore | imfindcircles