amd
Approximate minimum degree permutation
Syntax
P = amd(A)
P = amd(A,opts)
Description
P = amd(A) returns the
approximate minimum degree permutation vector for the sparse matrix C
= A + A'. The Cholesky factorization of C(P,P) or A(P,P) tends
to be sparser than that of C or A.
The amd function tends to be faster than symamd, and also tends to return better
orderings than symamd. Matrix A must
be square. If A is a full matrix, then amd(A) is
equivalent to amd(sparse(A)).
P = amd(A,opts) allows additional
options for the reordering. The opts input is a
structure with the two fields shown below. You only need to set the
fields of interest:
dense — A nonnegative scalar value that indicates what is considered to be dense. If A is n-by-n, then rows and columns with more than
max(16,(dense*sqrt(n)))entries inA + A'are considered to be "dense" and are ignored during the ordering. MATLAB® software places these rows and columns last in the output permutation. The default value for this field is 10.0 if this option is not present.aggressive — A scalar value controlling aggressive absorption. If this field is set to a nonzero value, then aggressive absorption is performed. This is the default if this option is not present.
MATLAB software performs an assembly tree post-ordering,
which is typically the same as an elimination tree post-ordering.
It is not always identical because of the approximate degree update
used, and because “dense” rows and columns do not take
part in the post-order. It well-suited for a subsequent chol operation, however, If you require
a precise elimination tree post-ordering, you can use the following
code:
P = amd(S); C = spones(S)+spones(S'); [ignore, Q] = etree(C(P,P)); P = P(Q);
If S is already symmetric, omit the second
line, C = spones(S)+spones(S').
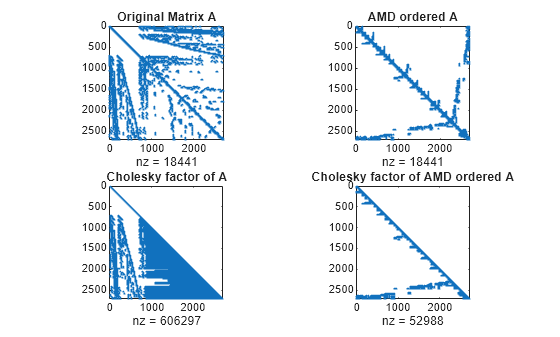
Examples
References
[1] Amestoy, Patrick R., Timothy A. Davis, and Iain S. Duff. “Algorithm 837: AMD, an Approximate Minimum Degree Ordering Algorithm.” ACM Transactions on Mathematical Software 30, no. 3 (September 2004): 381–388. https://doi.org/10.1145/1024074.1024081.