pyulear
Autoregressive power spectral density estimate — Yule-Walker method
Syntax
Description
pxx = pyulear(x,order)pxx, of
a discrete-time signal, x, found using the Yule-Walker
method. When x is a vector, it is treated as
a single channel. When x is a matrix, the PSD
is computed independently for each column and stored in the corresponding
column of pxx. pxx is the
distribution of power per unit frequency. The frequency is expressed
in units of rad/sample. order is the order of
the autoregressive (AR) model used to produce the PSD estimate.
pxx = pyulear(x,order,nfft)nfft points
in the discrete Fourier transform (DFT). For real x, pxx has
length (nfft/2 + 1)
if nfft is even, and (nfft + 1)/2 if nfft is
odd. For complex-valued x, pxx always
has length nfft. If you omit nfft,
or specify it as empty, then pyulear uses a default
DFT length of 256.
[
returns a frequency vector, pxx,f] = pyulear(___,fs)f, in cycles per unit time. The
sample rate, fs, is the number of samples per unit time. If
the unit of time is seconds, then f is in cycles/second
(Hz). For real–valued signals, f spans the interval
[0,fs/2] when nfft is even and
[0,fs/2) when nfft is odd. For
complex-valued signals, f spans the interval
[0,fs). fs must be the fourth
input to pyulear. To input a sample rate and still use
the default values of the preceding optional arguments, specify these arguments
as empty, [].
[
returns the two-sided AR PSD estimates at the frequencies specified in the
vector, pxx,f] = pyulear(x,order,f,fs)f. The vector, f, must contain
at least two elements, because otherwise the function interprets it as
nfft. The frequencies in f are in
cycles per unit time. The sample rate, fs, is the number of
samples per unit time. If the unit of time is seconds, then
f is in cycles/second (Hz).
[___, returns
the pxxc] = pyulear(___,'ConfidenceLevel',probability)probability × 100%
confidence intervals for the PSD estimate in pxxc.
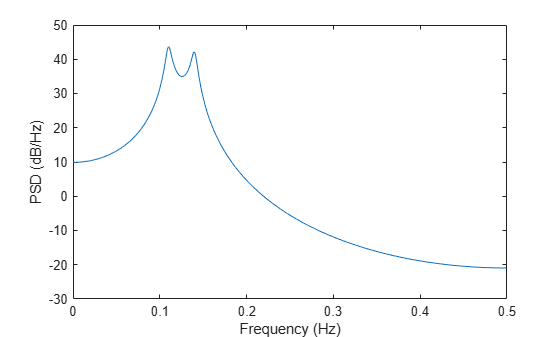
pyulear(___) with no output
arguments plots the AR PSD estimate in dB per unit frequency in the
current figure window.
Examples
Input Arguments
Output Arguments
Extended Capabilities
Version History
Introduced before R2006a