SimBiology Model Component Libraries
The SimBiology® libraries are collections of built-in kinetic laws, units, and unit prefixes that you can use while configuring reactions and quantity units in your model.
The built-in Abstract Kinetic Laws library provides a list of predefined reaction rates that follow particular kinetics, such as the mass action or Michaelis-Menten kinetics. The kinetic law for a newly added reaction is configured to MassAction by default. If you are using SimBiology Model Builder, the app automatically creates and maps the species and parameters needed by the reaction rate. For other kinetic laws, only parameters are created and mapped. You need to create and map the species manually. Use the Unknown kinetic law to define a custom reaction rate with its own parameters. You must define and add the species and parameters needed by the custom rate.
Note
The MassAction and Unknown kinetic laws can have different simulation results even when the reaction rate is the same. This can happen when you have a reversible reaction with species in different compartments. The difference in simulation results is because of the volume-scaling performed by SimBiology during the dimensional analysis. For details, see Derive ODEs from SimBiology Reactions. Specifically, for MassAction, SimBiology uses corresponding compartment volumes to multiply the forward and reverse rates. However, for Unknown and other built-in kinetic laws, SimBiology multiplies the entire rate by only one compartment which contains the reactants. To see exactly what compartment volumes are used for scaling, use getequations or open the Equations view from the SimBiology app and check the ODEs section.
The Units library provides a list of available units and corresponding unit compositions. The Unit Prefixes library provides a list of prefixes and corresponding exponent values for unit prefixes.
You can also add custom components to any library. For instance, you can define a custom unit and use it in your model. These custom components are saved across sessions of MATLAB®. The libraries are available for all SimBiology projects and are not part of any one project.
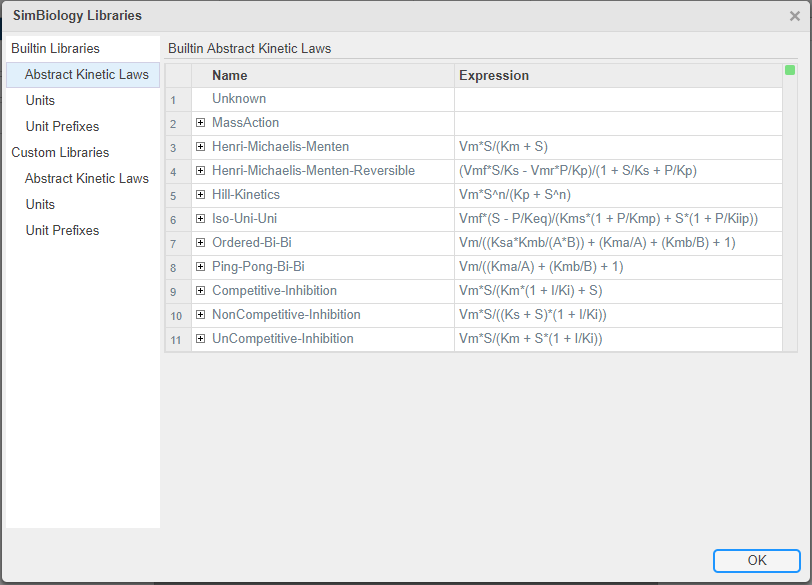
To see the list of libraries in the SimBiology Model Builder app, click Libraries on the Home tab. The following figure shows the kinetic laws library with all available built-in kinetic laws.
See Also
SimBiology Model Builder | SimBiology Model Analyzer