BayesianOptimization
Bayesian optimization results
Description
A BayesianOptimization object contains the
results of a Bayesian optimization. It is the output of bayesopt or a fit function that accepts the
OptimizeHyperparameters name-value pair such as fitcdiscr. In addition, a BayesianOptimization object contains data for each iteration of
bayesopt that can be accessed by a plot function or an output
function.
Creation
Create a BayesianOptimization object by using the
bayesopt function or one of the following
fit functions with the OptimizeHyperparameters name-value
argument.
Classification fit functions:
fitcdiscr,fitcecoc,fitcensemble,fitcgam,fitckernel,fitcknn,fitclinear,fitcnb,fitcnet,fitcsvm,fitctreeRegression fit functions:
fitrensemble,fitrgam,fitrgp,fitrkernel,fitrlinear,fitrnet,fitrsvm,fitrtree
Properties
Problem Definition Properties
ObjectiveFcn — ObjectiveFcn argument used by
bayesopt
function handle
This property is read-only.
ObjectiveFcn argument used by
bayesopt, specified as a function handle.
If you call
bayesoptdirectly,ObjectiveFcnis thebayesoptobjective function argument.If you call a fit function containing the
'OptimizeHyperparameters'name-value pair argument,ObjectiveFcnis a function handle that returns the misclassification rate for classification or returns the logarithm of one plus the cross-validation loss for regression, measured by five-fold cross-validation.
Data Types: function_handle
VariableDescriptions — VariableDescriptions argument that bayesopt used
vector of optimizableVariable objects
This property is read-only.
VariableDescriptions argument that
bayesopt used, specified as a vector of optimizableVariable
objects.
If you called
bayesoptdirectly,VariableDescriptionsis thebayesoptvariable description argument.If you called a fit function with the
OptimizeHyperparametersname-value pair,VariableDescriptionsis the vector of hyperparameters.
Options — Options that bayesopt used
structure
This property is read-only.
Options that bayesopt used, specified as a
structure.
If you called
bayesoptdirectly,Optionsis the options used inbayesopt, which are the name-value pairs SeebayesoptInput Arguments.If you called a fit function with the
OptimizeHyperparametersname-value pair,Optionsare the defaultbayesoptoptions, modified by theHyperparameterOptimizationOptionsname-value pair.
Options is a read-only structure containing the
following fields.
| Option Name | Meaning |
|---|---|
AcquisitionFunctionName | Acquisition function name. See Acquisition Function Types. |
IsObjectiveDeterministic | true means the objective function
is deterministic, false
otherwise. |
ExplorationRatio | Used only when
AcquisitionFunctionName is
'expected-improvement-plus' or
'expected-improvement-per-second-plus'.
See Plus. |
MaxObjectiveEvaluations | Objective function evaluation limit. |
MaxTime | Time limit. |
XConstraintFcn | Deterministic constraints on variables. See Deterministic Constraints — XConstraintFcn. |
ConditionalVariableFcn | Conditional variable constraints. See Conditional Constraints — ConditionalVariableFcn. |
NumCoupledConstraints | Number of coupled constraints. See Coupled Constraints. |
CoupledConstraintTolerances | Coupled constraint tolerances. See Coupled Constraints. |
AreCoupledConstraintsDeterministic | Logical vector specifying whether each coupled constraint is deterministic. |
Verbose | Command-line display level. |
OutputFcn | Function called after each iteration. See Bayesian Optimization Output Functions. |
SaveVariableName | Variable name for the
@assignInBase output function.
|
SaveFileName | File name for the @saveToFile
output function. |
PlotFcn | Plot function called after each iteration. See Bayesian Optimization Plot Functions |
InitialX | Points where bayesopt evaluated
the objective function. |
InitialObjective | Objective function values at
InitialX. |
InitialConstraintViolations | Coupled constraint function values at
InitialX. |
InitialErrorValues | Error values at InitialX. |
InitialObjectiveEvaluationTimes | Objective function evaluation times at
InitialX. |
InitialIterationTimes | Time for each iteration, including objective function evaluation and other computations. |
Data Types: struct
Solution Properties
MinObjective — Minimum observed value of objective function
real scalar
This property is read-only.
Minimum observed value of objective function, specified as a real scalar. When there are coupled constraints or evaluation errors, this value is the minimum over all observed points that are feasible according to the final constraint and Error models.
Data Types: double
XAtMinObjective — Observed point with minimum objective function value
1-by-D table
This property is read-only.
Observed point with minimum objective function value, specified as a
1-by-D table, where
D is the number of variables.
Data Types: table
MinEstimatedObjective — Estimated objective function value
real scalar
This property is read-only.
Estimated objective function value at
XAtMinEstimatedObjective, specified as a real
scalar.
MinEstimatedObjective is the mean value of the
posterior distribution of the final objective model. The software
estimates the MinEstimatedObjective value by
passing XAtMinEstimatedObjective to the object
function predictObjective.
Data Types: double
XAtMinEstimatedObjective — Point with minimum upper confidence bound of objective function value
1-by-D table
This property is read-only.
Point with the minimum upper confidence bound of the objective
function value among the visited points, specified as a
1-by-D table, where
D is the number of variables. The software uses
the final objective model to find the upper confidence bounds of the
visited points.
XAtMinEstimatedObjective is the same as the best
point returned by the bestPoint function with the
default criterion
('min-visited-upper-confidence-interval').
Data Types: table
NumObjectiveEvaluations — Number of objective function evaluations
positive integer
This property is read-only.
Number of objective function evaluations, specified as a positive integer. This includes the initial evaluations to form a posterior model as well as evaluation during the optimization iterations.
Data Types: double
TotalElapsedTime — Total elapsed time of optimization in seconds
positive scalar
This property is read-only.
Total elapsed time of optimization in seconds, specified as a positive scalar.
Data Types: double
NextPoint — Next point to evaluate if optimization continues
1-by-D table
This property is read-only.
Next point to evaluate if optimization continues, specified as a
1-by-D table, where
D is the number of variables.
Data Types: table
Trace Properties
XTrace — Points where the objective function was evaluated
T-by-D table
This property is read-only.
Points where the objective function was evaluated, specified as a
T-by-D table, where
T is the number of evaluation points and
D is the number of variables.
Data Types: table
ObjectiveTrace — Objective function values
column vector of length T
This property is read-only.
Objective function values, specified as a column vector of length
T, where T is the number of
evaluation points. ObjectiveTrace contains the
history of objective function evaluations.
Data Types: double
ObjectiveEvaluationTimeTrace — Objective function evaluation times
column vector of length T
This property is read-only.
Objective function evaluation times, specified as a column vector of
length T, where T is the number of
evaluation points. ObjectiveEvaluationTimeTrace
includes the time in evaluating coupled constraints, because the
objective function computes these constraints.
Data Types: double
IterationTimeTrace — Iteration times
column vector of length T
This property is read-only.
Iteration times, specified as a column vector of length
T, where T is the number of
evaluation points. IterationTimeTrace includes both
objective function evaluation time and other overhead.
Data Types: double
ConstraintsTrace — Coupled constraint values
T-by-K array
This property is read-only.
Coupled constraint values, specified as a
T-by-K array, where
T is the number of evaluation points and
K is the number of coupled constraints.
Data Types: double
ErrorTrace — Error indications
column vector of length T of -1
or 1 entries
This property is read-only.
Error indications, specified as a column vector of length
T of -1 or
1 entries, where T is the
number of evaluation points. Each 1 entry indicates
that the objective function errored or returned NaN
on the corresponding point in XTrace. Each
-1 entry indicates that the objective function
value was computed.
Data Types: double
FeasibilityTrace — Feasibility indications
logical column vector of length T
This property is read-only.
Feasibility indications, specified as a logical column vector of
length T, where T is the number of
evaluation points. Each 1 entry indicates that the
final constraint model predicts feasibility at the corresponding point
in XTrace.
Data Types: logical
FeasibilityProbabilityTrace — Probability that evaluation point is feasible
column vector of length T
This property is read-only.
Probability that evaluation point is feasible, specified as a column
vector of length T, where T is the
number of evaluation points. The probabilities come from the final
constraint model, including the error constraint model, on the
corresponding points in XTrace.
Data Types: double
IndexOfMinimumTrace — Which evaluation gave minimum feasible objective
column vector of integer indices of length
T
This property is read-only.
Which evaluation gave minimum feasible objective, specified as a
column vector of integer indices of length T, where
T is the number of evaluation points. Feasibility
is determined with respect to the constraint models that existed at each
iteration, including the error constraint model.
Data Types: double
ObjectiveMinimumTrace — Minimum observed objective
column vector of length T
This property is read-only.
Minimum observed objective, specified as a column vector of length
T, where T is the number of
evaluation points.
Data Types: double
EstimatedObjectiveMinimumTrace — Estimated objective
column vector of length T
This property is read-only.
Estimated objective, specified as a column vector of length
T, where T is the number of
evaluation points. The estimated objective at each iteration is
determined with respect to the objective model at that iteration. At
each iteration, the software uses the object function predictObjective to
estimate the objective function value at the point with the minimum
upper confidence bound of the objective function among the visited
points.
Data Types: double
UserDataTrace — Auxiliary data from the objective function
cell array of length T
This property is read-only.
Auxiliary data from the objective function, specified as a cell array
of length T, where T is the number
of evaluation points. Each entry in the cell array is the
UserData returned in the third output of the
objective function.
Data Types: cell
Object Functions
bestPoint | Best point in a Bayesian optimization according to a criterion |
plot | Plot Bayesian optimization results |
predictConstraints | Predict coupled constraint violations at a set of points |
predictError | Predict error value at a set of points |
predictObjective | Predict objective function at a set of points |
predictObjectiveEvaluationTime | Predict objective function run times at a set of points |
resume | Resume a Bayesian optimization |
Examples
Create a BayesianOptimization Object Using bayesopt
This example shows how to create a BayesianOptimization object by using bayesopt to minimize cross-validation loss.
Optimize hyperparameters of a KNN classifier for the ionosphere data, that is, find KNN hyperparameters that minimize the cross-validation loss. Have bayesopt minimize over the following hyperparameters:
Nearest-neighborhood sizes from 1 to 30
Distance functions
'chebychev','euclidean', and'minkowski'.
For reproducibility, set the random seed, set the partition, and set the AcquisitionFunctionName option to 'expected-improvement-plus'. To suppress iterative display, set 'Verbose' to 0. Pass the partition c and fitting data X and Y to the objective function fun by creating fun as an anonymous function that incorporates this data. See Parameterizing Functions.
load ionosphere rng default num = optimizableVariable('n',[1,30],'Type','integer'); dst = optimizableVariable('dst',{'chebychev','euclidean','minkowski'},'Type','categorical'); c = cvpartition(351,'Kfold',5); fun = @(x)kfoldLoss(fitcknn(X,Y,'CVPartition',c,'NumNeighbors',x.n,... 'Distance',char(x.dst),'NSMethod','exhaustive')); results = bayesopt(fun,[num,dst],'Verbose',0,... 'AcquisitionFunctionName','expected-improvement-plus')
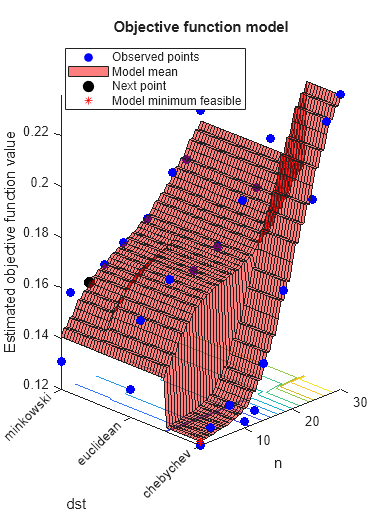
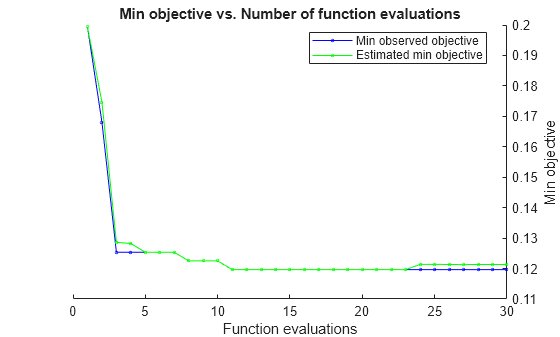
results =
BayesianOptimization with properties:
ObjectiveFcn: @(x)kfoldLoss(fitcknn(X,Y,'CVPartition',c,'NumNeighbors',x.n,'Distance',char(x.dst),'NSMethod','exhaustive'))
VariableDescriptions: [1x2 optimizableVariable]
Options: [1x1 struct]
MinObjective: 0.1197
XAtMinObjective: [1x2 table]
MinEstimatedObjective: 0.1213
XAtMinEstimatedObjective: [1x2 table]
NumObjectiveEvaluations: 30
TotalElapsedTime: 36.7111
NextPoint: [1x2 table]
XTrace: [30x2 table]
ObjectiveTrace: [30x1 double]
ConstraintsTrace: []
UserDataTrace: {30x1 cell}
ObjectiveEvaluationTimeTrace: [30x1 double]
IterationTimeTrace: [30x1 double]
ErrorTrace: [30x1 double]
FeasibilityTrace: [30x1 logical]
FeasibilityProbabilityTrace: [30x1 double]
IndexOfMinimumTrace: [30x1 double]
ObjectiveMinimumTrace: [30x1 double]
EstimatedObjectiveMinimumTrace: [30x1 double]
Create a BayesianOptimization Object Using a Fit Function
This example shows how to minimize the cross-validation loss in the ionosphere data using Bayesian optimization of an SVM classifier.
Load the data.
load ionosphereOptimize the classification using the 'auto' parameters.
rng default % For reproducibility Mdl = fitcsvm(X,Y,'OptimizeHyperparameters','auto')
|====================================================================================================================|
| Iter | Eval | Objective | Objective | BestSoFar | BestSoFar | BoxConstraint| KernelScale | Standardize |
| | result | | runtime | (observed) | (estim.) | | | |
|====================================================================================================================|
| 1 | Best | 0.35897 | 0.40705 | 0.35897 | 0.35897 | 3.8653 | 961.53 | true |
| 2 | Best | 0.12821 | 9.5202 | 0.12821 | 0.15646 | 429.99 | 0.2378 | false |
| 3 | Accept | 0.35897 | 0.20668 | 0.12821 | 0.1315 | 0.11801 | 8.9479 | false |
| 4 | Accept | 0.1339 | 4.6054 | 0.12821 | 0.12965 | 0.0010694 | 0.0032063 | true |
| 5 | Accept | 0.15954 | 10.71 | 0.12821 | 0.12824 | 973.65 | 0.15179 | false |
| 6 | Accept | 0.35897 | 0.064195 | 0.12821 | 0.12826 | 0.0010106 | 146.16 | false |
| 7 | Best | 0.12536 | 0.18824 | 0.12536 | 0.14287 | 0.00102 | 0.016781 | false |
| 8 | Accept | 0.12821 | 0.14486 | 0.12536 | 0.12794 | 0.0010081 | 0.045557 | false |
| 9 | Best | 0.12251 | 0.097958 | 0.12251 | 0.12511 | 0.0010204 | 0.042244 | false |
| 10 | Accept | 0.13675 | 0.30522 | 0.12251 | 0.12547 | 0.0064447 | 0.022499 | false |
| 11 | Accept | 0.16809 | 0.17466 | 0.12251 | 0.12514 | 0.0010017 | 0.33772 | false |
| 12 | Accept | 0.1339 | 5.7734 | 0.12251 | 0.12378 | 0.0010063 | 0.0015104 | false |
| 13 | Accept | 0.21368 | 0.074162 | 0.12251 | 0.12383 | 140.53 | 151.78 | false |
| 14 | Accept | 0.35897 | 0.060885 | 0.12251 | 0.12579 | 0.0010305 | 297.99 | true |
| 15 | Accept | 0.12536 | 0.08714 | 0.12251 | 0.12568 | 992.34 | 21.453 | false |
| 16 | Accept | 0.13105 | 0.19838 | 0.12251 | 0.12588 | 386.2 | 6.3489 | false |
| 17 | Accept | 0.14245 | 0.16694 | 0.12251 | 0.12358 | 998.93 | 7.0729 | false |
| 18 | Accept | 0.1396 | 0.19816 | 0.12251 | 0.12346 | 996.24 | 211.36 | false |
| 19 | Accept | 0.12251 | 0.18788 | 0.12251 | 0.1229 | 0.0020931 | 0.025996 | false |
| 20 | Accept | 0.1339 | 5.8506 | 0.12251 | 0.12293 | 972.08 | 1.7749 | true |
|====================================================================================================================|
| Iter | Eval | Objective | Objective | BestSoFar | BestSoFar | BoxConstraint| KernelScale | Standardize |
| | result | | runtime | (observed) | (estim.) | | | |
|====================================================================================================================|
| 21 | Accept | 0.20228 | 10.503 | 0.12251 | 0.12284 | 995.93 | 0.0012925 | true |
| 22 | Accept | 0.151 | 0.12298 | 0.12251 | 0.12267 | 970.91 | 623.29 | true |
| 23 | Best | 0.11966 | 0.093049 | 0.11966 | 0.12022 | 983.19 | 63.935 | true |
| 24 | Best | 0.11681 | 0.062309 | 0.11681 | 0.11735 | 343.06 | 64.188 | true |
| 25 | Accept | 0.12821 | 0.19236 | 0.11681 | 0.1224 | 456.25 | 28.864 | true |
| 26 | Accept | 0.11966 | 0.068103 | 0.11681 | 0.11954 | 477.93 | 99.229 | true |
| 27 | Accept | 0.1339 | 0.11542 | 0.11681 | 0.1199 | 225.77 | 149.38 | true |
| 28 | Accept | 0.17094 | 10.6 | 0.11681 | 0.12001 | 20.868 | 0.010729 | false |
| 29 | Accept | 0.11681 | 0.12386 | 0.11681 | 0.11776 | 628.64 | 69.451 | true |
| 30 | Accept | 0.1396 | 0.30821 | 0.11681 | 0.11776 | 82.05 | 2.3607 | false |
__________________________________________________________
Optimization completed.
MaxObjectiveEvaluations of 30 reached.
Total function evaluations: 30
Total elapsed time: 75.6701 seconds
Total objective function evaluation time: 61.2117
Best observed feasible point:
BoxConstraint KernelScale Standardize
_____________ ___________ ___________
343.06 64.188 true
Observed objective function value = 0.11681
Estimated objective function value = 0.12068
Function evaluation time = 0.062309
Best estimated feasible point (according to models):
BoxConstraint KernelScale Standardize
_____________ ___________ ___________
628.64 69.451 true
Estimated objective function value = 0.11776
Estimated function evaluation time = 0.091281
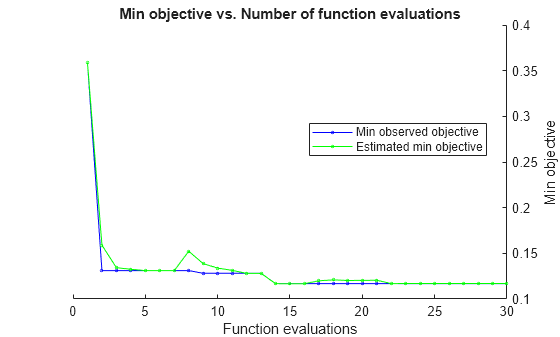
Mdl =
ClassificationSVM
ResponseName: 'Y'
CategoricalPredictors: []
ClassNames: {'b' 'g'}
ScoreTransform: 'none'
NumObservations: 351
HyperparameterOptimizationResults: [1x1 BayesianOptimization]
Alpha: [110x1 double]
Bias: 0.2184
KernelParameters: [1x1 struct]
Mu: [0.8917 0 0.6413 0.0444 0.6011 0.1159 0.5501 0.1194 0.5118 0.1813 0.4762 0.1550 0.4008 0.0934 0.3442 0.0711 0.3819 -0.0036 0.3594 -0.0240 0.3367 0.0083 0.3625 -0.0574 0.3961 -0.0712 0.5416 -0.0695 ... ] (1x34 double)
Sigma: [0.3112 0 0.4977 0.4414 0.5199 0.4608 0.4927 0.5207 0.5071 0.4839 0.5635 0.4948 0.6222 0.4949 0.6528 0.4584 0.6180 0.4968 0.6263 0.5191 0.6098 0.5182 0.6038 0.5275 0.5785 0.5085 0.5162 0.5500 ... ] (1x34 double)
BoxConstraints: [351x1 double]
ConvergenceInfo: [1x1 struct]
IsSupportVector: [351x1 logical]
Solver: 'SMO'
The fit achieved about 12% loss for the default 5-fold cross validation.
Examine the BayesianOptimization object that is returned in the HyperparameterOptimizationResults property of the returned model.
disp(Mdl.HyperparameterOptimizationResults)
BayesianOptimization with properties:
ObjectiveFcn: @createObjFcn/inMemoryObjFcn
VariableDescriptions: [5x1 optimizableVariable]
Options: [1x1 struct]
MinObjective: 0.1168
XAtMinObjective: [1x3 table]
MinEstimatedObjective: 0.1178
XAtMinEstimatedObjective: [1x3 table]
NumObjectiveEvaluations: 30
TotalElapsedTime: 75.6701
NextPoint: [1x3 table]
XTrace: [30x3 table]
ObjectiveTrace: [30x1 double]
ConstraintsTrace: []
UserDataTrace: {30x1 cell}
ObjectiveEvaluationTimeTrace: [30x1 double]
IterationTimeTrace: [30x1 double]
ErrorTrace: [30x1 double]
FeasibilityTrace: [30x1 logical]
FeasibilityProbabilityTrace: [30x1 double]
IndexOfMinimumTrace: [30x1 double]
ObjectiveMinimumTrace: [30x1 double]
EstimatedObjectiveMinimumTrace: [30x1 double]
Version History
Introduced in R2016b
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.

Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list:
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)