confusionmat
Compute confusion matrix for classification problem
Syntax
Description
C = confusionmat(group,grouphat,'Order',grouporder)grouporder to order the rows and columns of C.
Examples
Display Confusion Matrix
Display the confusion matrix for data with two misclassifications and one missing classification.
Create vectors for the known groups and the predicted groups.
g1 = [3 2 2 3 1 1]'; % Known groups g2 = [4 2 3 NaN 1 1]'; % Predicted groups
Return the confusion matrix.
C = confusionmat(g1,g2)
C = 4×4
2 0 0 0
0 1 1 0
0 0 0 1
0 0 0 0
The indices of the rows and columns of the confusion matrix C are identical and arranged by default in the sorted order of [g1;g2], that is, (1,2,3,4).
The confusion matrix shows that the two data points known to be in group 1 are classified correctly. For group 2, one of the data points is misclassified into group 3. Also, one of the data points known to be in group 3 is misclassified into group 4. confusionmat treats the NaN value in the grouping variable g2 as a missing value and does not include it in the rows and columns of C.
Plot the confusion matrix as a confusion matrix chart by using confusionchart.
confusionchart(C)
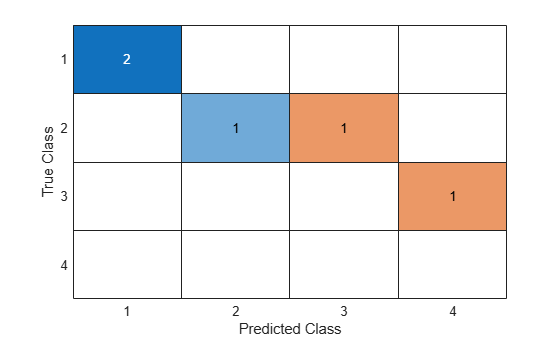
You do not need to calculate the confusion matrix first and then plot it. Instead, plot a confusion matrix chart directly from the true and predicted labels by using confusionchart.
cm = confusionchart(g1,g2)
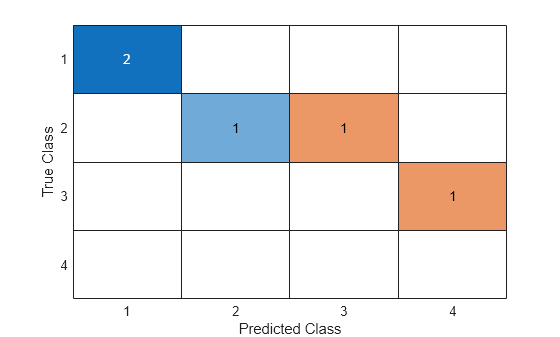
cm =
ConfusionMatrixChart with properties:
NormalizedValues: [4x4 double]
ClassLabels: [4x1 double]
Use GET to show all properties
The ConfusionMatrixChart object stores the numeric confusion matrix in the NormalizedValues property and the classes in the ClassLabels property. Display these properties using dot notation.
cm.NormalizedValues
ans = 4×4
2 0 0 0
0 1 1 0
0 0 0 1
0 0 0 0
cm.ClassLabels
ans = 4×1
1
2
3
4
Specify Group Order of Confusion Matrix
Display the confusion matrix for data with two misclassifications and one missing classification, and specify the group order.
Create vectors for the known groups and the predicted groups.
g1 = [3 2 2 3 1 1]'; % Known groups g2 = [4 2 3 NaN 1 1]'; % Predicted groups
Specify the group order and return the confusion matrix.
C = confusionmat(g1,g2,'Order',[4 3 2 1])C = 4×4
0 0 0 0
1 0 0 0
0 1 1 0
0 0 0 2
The indices of the rows and columns of the confusion matrix C are identical and arranged in the order specified by the group order, that is, (4,3,2,1).
The second row of the confusion matrix C shows that one of the data points known to be in group 3 is misclassified into group 4. The third row of C shows that one of the data points belonging to group 2 is misclassified into group 3, and the fourth row shows that the two data points known to be in group 1 are classified correctly. confusionmat treats the NaN value in the grouping variable g2 as a missing value and does not include it in the rows and columns of C.
Confusion Matrix for Classification
Perform classification on a sample of the fisheriris data set and display the confusion matrix for the resulting classification.
Load Fisher's iris data set.
load fisheririsRandomize the measurements and groups in the data.
rng(0,'twister'); % For reproducibility numObs = length(species); p = randperm(numObs); meas = meas(p,:); species = species(p);
Train a discriminant analysis classifier by using measurements in the first half of the data.
half = floor(numObs/2); training = meas(1:half,:); trainingSpecies = species(1:half); Mdl = fitcdiscr(training,trainingSpecies);
Predict labels for the measurements in the second half of the data by using the trained classifier.
sample = meas(half+1:end,:); grouphat = predict(Mdl,sample);
Specify the group order and display the confusion matrix for the resulting classification.
group = species(half+1:end); [C,order] = confusionmat(group,grouphat,'Order',{'setosa','versicolor','virginica'})
C = 3×3
29 0 0
0 22 2
0 0 22
order = 3x1 cell
{'setosa' }
{'versicolor'}
{'virginica' }
The confusion matrix shows that the measurements belonging to setosa and virginica are classified correctly, while two of the measurements belonging to versicolor are misclassified as virginica. The output order contains the order of the rows and columns of the confusion matrix in the sequence specified by the group order {'setosa','versicolor','virginica'}.
Confusion Matrix for Classification Using Tall Arrays
Perform classification on a tall array of the fisheriris data set, compute a confusion matrix for the known and predicted tall labels by using the confusionmat function, and plot the confusion matrix by using the confusionchart function.
When you perform calculations on tall arrays, MATLAB® uses either a parallel pool (default if you have Parallel Computing Toolbox™) or the local MATLAB session. If you want to run the example using the local MATLAB session when you have Parallel Computing Toolbox, you can change the global execution environment by using the mapreducer function.
Load Fisher's iris data set.
load fisheririsConvert the in-memory arrays meas and species to tall arrays.
tx = tall(meas);
Starting parallel pool (parpool) using the 'local' profile ... Connected to the parallel pool (number of workers: 6).
ty = tall(species);
Find the number of observations in the tall array.
numObs = gather(length(ty)); % gather collects tall array into memorySet the seeds of the random number generators using rng and tallrng for reproducibility, and randomly select training samples. The results can vary depending on the number of workers and the execution environment for the tall arrays. For details, see Control Where Your Code Runs.
rng('default') tallrng('default') numTrain = floor(numObs/2); [txTrain,trIdx] = datasample(tx,numTrain,'Replace',false); tyTrain = ty(trIdx);
Fit a decision tree classifier model on the training samples.
mdl = fitctree(txTrain,tyTrain);
Evaluating tall expression using the Parallel Pool 'local': - Pass 1 of 2: Completed in 3.9 sec - Pass 2 of 2: Completed in 1.5 sec Evaluation completed in 7.3 sec Evaluating tall expression using the Parallel Pool 'local': - Pass 1 of 4: Completed in 0.88 sec - Pass 2 of 4: Completed in 1.6 sec - Pass 3 of 4: Completed in 4 sec - Pass 4 of 4: Completed in 2.7 sec Evaluation completed in 11 sec Evaluating tall expression using the Parallel Pool 'local': - Pass 1 of 4: Completed in 0.54 sec - Pass 2 of 4: Completed in 1.2 sec - Pass 3 of 4: Completed in 3 sec - Pass 4 of 4: Completed in 2 sec Evaluation completed in 7.6 sec Evaluating tall expression using the Parallel Pool 'local': - Pass 1 of 4: Completed in 0.51 sec - Pass 2 of 4: Completed in 1.3 sec - Pass 3 of 4: Completed in 3.1 sec - Pass 4 of 4: Completed in 2.5 sec Evaluation completed in 8.5 sec Evaluating tall expression using the Parallel Pool 'local': - Pass 1 of 4: Completed in 0.42 sec - Pass 2 of 4: Completed in 1.2 sec - Pass 3 of 4: Completed in 3 sec - Pass 4 of 4: Completed in 2.1 sec Evaluation completed in 7.6 sec
Predict labels for the test samples by using the trained model.
txTest = tx(~trIdx,:); label = predict(mdl,txTest);
Compute the confusion matrix for the resulting classification.
tyTest = ty(~trIdx); [C,order] = confusionmat(tyTest,label)
C =
M×N×... tall array
? ? ? ...
? ? ? ...
? ? ? ...
: : :
: : :
Preview deferred. Learn more.
order =
M×N×... tall array
? ? ? ...
? ? ? ...
? ? ? ...
: : :
: : :
Preview deferred. Learn more.
Use the gather function to perform the deferred calculation and return the result of confusionmat in memory.
gather(C)
Evaluating tall expression using the Parallel Pool 'local': - Pass 1 of 1: Completed in 1.9 sec Evaluation completed in 2.3 sec
ans = 3×3
20 0 0
1 30 2
0 0 22
gather(order)
Evaluating tall expression using the Parallel Pool 'local': Evaluation completed in 0.032 sec
ans = 3×1 cell
{'setosa' }
{'versicolor'}
{'virginica' }
The confusion matrix shows that three measurements in the versicolor class are misclassified. All the measurements belonging to setosa and virginica are classified correctly.
To compute and plot the confusion matrix, use confusionchart instead.
cm = confusionchart(tyTest,label)
Evaluating tall expression using the Parallel Pool 'local': - Pass 1 of 1: Completed in 0.34 sec Evaluation completed in 0.6 sec Evaluating tall expression using the Parallel Pool 'local': - Pass 1 of 1: Completed in 0.48 sec Evaluation completed in 0.67 sec
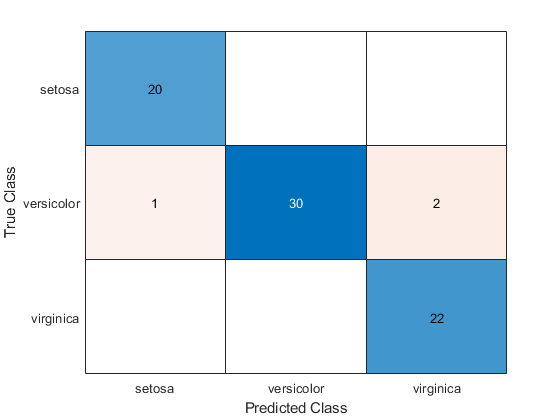
cm =
ConfusionMatrixChart with properties:
NormalizedValues: [3×3 double]
ClassLabels: {3×1 cell}
Show all properties
Display Matrix of Frequencies
Use the confusionmat function to create a matrix showing the number of flights that travel between airports listed in the columns of a tall table.
When you perform calculations on tall arrays, MATLAB® uses either a parallel pool (default if you have Parallel Computing Toolbox™) or the local MATLAB session. To run the example using the local MATLAB session when you have Parallel Computing Toolbox, change the global execution environment by using the mapreducer function.
mapreducer(0)
Create a datastore for the airlinesmall.csv data set. Treat 'NA' values as missing data so that they are replaced with NaN values. Select the variables Origin and Dest to include in the datastore.
varnames = {'Origin','Dest'};
ds = datastore('airlinesmall.csv','TreatAsMissing','NA', ...
'SelectedVariableNames',varnames);Create a tall array for the data in the datastore. Because the data in ds is tabular, the result is a tall table. If the data is not tabular, then tall creates a tall cell array instead.
T = tall(ds)
T =
Mx2 tall table
Origin Dest
_______ _______
{'LAX'} {'SJC'}
{'SJC'} {'BUR'}
{'SAN'} {'SMF'}
{'BUR'} {'SJC'}
{'SMF'} {'LAX'}
{'LAX'} {'SJC'}
{'SAN'} {'SFO'}
{'SEA'} {'LAX'}
: :
: :
The display of the tall table indicates that the number of rows of data is unknown.
Create a matrix showing the number of flights between columns T.Origin and T.Dest. This matrix is not a confusion matrix, because the two columns do not contain known and predicted values from classification. However, you can use the confusionmat function to create a matrix of frequencies.
[ta,tb] = confusionmat(T.Origin,T.Dest)
ta =
MxNx... tall array
? ? ? ...
? ? ? ...
? ? ? ...
: : :
: : :
tb =
MxNx... tall array
? ? ? ...
? ? ? ...
? ? ? ...
: : :
: : :
Perform the deferred calculation by using the gather function, and return the result of confusionmat in memory.
[freqMatrix,airportOrder] = gather(ta,tb);
Evaluating tall expression using the Local MATLAB Session: - Pass 1 of 1: Completed in 1.3 sec Evaluation completed in 1.7 sec
Display the first five rows of the matrix freqMatrix and the corresponding order of rows and columns airportOrder.
freqMatrix(1:5,:)
ans = 5×323
0 153 169 0 91 161 322 0 44 6 56 24 0 0 23 180 122 20 150 20 63 77 134 37 10 0 3 51 0 1 311 0 15 0 32 81 30 53 0 9 2 15 12 293 20 38 1 73 0 41
168 0 75 59 5 76 0 6 14 79 0 1 0 0 0 54 60 0 5 0 1 5 51 0 0 0 0 1 0 0 55 0 0 0 8 67 50 0 0 0 0 18 1 59 1 0 0 11 0 4
187 87 0 0 78 39 120 0 14 1 18 19 0 0 0 98 95 2 19 3 14 14 72 0 0 0 0 0 0 0 108 0 1 0 1 31 4 14 0 1 0 3 9 172 5 13 0 21 0 10
0 58 0 0 61 25 83 3 2 1 0 0 0 0 0 0 23 0 5 0 0 0 21 0 0 0 0 0 0 0 87 0 0 0 0 13 0 0 0 0 0 0 0 67 0 0 0 1 0 0
114 1 88 73 0 70 20 5 4 47 1 3 0 0 0 40 39 0 1 0 0 3 57 0 0 0 0 0 0 0 50 0 1 0 1 28 1 0 0 0 0 0 2 58 5 0 0 21 0 0
airportOrder(1:5)
ans = 5x1 cell
{'LAX'}
{'SJC'}
{'SAN'}
{'BUR'}
{'SMF'}
The matrix freqMatrix displays the number of flights from an origin airport (row) to a destination airport (column). For example, a total of 168 flights leave SJC and arrive at LAX (see freqMatrix(2,1)). Similarly, 88 flights leave SMF and arrive at SAN (see freqMatrix(5,3)). As noted earlier, freqMatrix is not a confusion matrix, but shows a count of flights between airports. As expected, the diagonal elements are all zeros, because the origin and destination airport are always different.
Input Arguments
group — Known groups
numeric vector | logical vector | character array | string array | cell array of character vectors | categorical vector
Known groups for categorizing observations, specified as a numeric vector, logical vector, character array, string array, cell array of character vectors, or categorical vector.
group is a grouping variable of the same type as grouphat. The group argument must have the same number of observations as grouphat, as described in Grouping Variables. The confusionmat function treats character arrays and string arrays as cell arrays of character vectors. Additionally, confusionmat treats NaN, empty, and 'undefined' values in group as missing values and does not count them as distinct groups or categories.
Example: {'Male','Female','Female','Male','Female'}
Data Types: single | double | logical | char | string | cell | categorical
grouphat — Predicted groups
numeric vector | logical vector | character array | string array | cell array of character vectors | categorical vector
Predicted groups for categorizing observations, specified as a numeric vector, logical vector, character array, string array, cell array of character vectors, or categorical vector.
grouphat is a grouping variable of the same type as group. The grouphat argument must have the same number of observations as group, as described in Grouping Variables. The confusionmat function treats character arrays and string arrays as cell arrays of character vectors. Additionally, confusionmat treats NaN, empty, and 'undefined' values in grouphat as missing values and does not count them as distinct groups or categories.
Example: [1 0 0 1 0]
Data Types: single | double | logical | char | string | cell | categorical
grouporder — Group order
numeric vector | logical vector | character array | string array | cell array of character vectors | categorical vector
Group order, specified as a numeric vector, logical vector, character array, string array, cell array of character vectors, or categorical vector.
grouporder is a grouping variable containing all the distinct elements in group and grouphat. Specify grouporder to define the order of the rows and columns of C. If grouporder contains elements that are not in group or grouphat, the corresponding entries in C are 0.
By default, the group order depends on the data type of s = [group;grouphat]:
For numeric and logical vectors, the order is the sorted order of
s.For categorical vectors, the order is the order returned by
categories(s)For other data types, the order is the order of first appearance in
s.
Example: 'order',{'setosa','versicolor','virginica'}
Data Types: single | double | logical | char | string | cell | categorical
Output Arguments
C — Confusion matrix
matrix
Confusion matrix, returned as a square matrix with size equal to the total number of distinct elements in the group and grouphat arguments. C(i,j) is the count of observations known to be in group i but predicted to be in group j.
The rows and columns of C have identical ordering of the same group indices. By default, the group order depends on the data type of s = [group;grouphat]:
For numeric and logical vectors, the order is the sorted order of
s.For categorical vectors, the order is the order returned by
categories(s)For other data types, the order is the order of first appearance in
s.
To change the order, specify grouporder,
The confusionmat function treats NaN, empty, and 'undefined' values in the grouping variables as missing values and does not include them in the rows and columns of C.
order — Order of rows and columns
numeric vector | logical vector | categorical vector | cell array of character vectors
Order of rows and columns in C, returned as a numeric vector, logical vector, categorical vector, or cell array of character vectors. If group and grouphat are character arrays, string arrays, or cell arrays of character vectors, then the variable order is a cell array of character vectors. Otherwise, order is of the same type as group and grouphat.
Alternative Functionality
Use
confusionchartto calculate and plot a confusion matrix. Additionally,confusionchartdisplays summary statistics about your data and sorts the classes of the confusion matrix according to the class-wise precision (positive predictive value), class-wise recall (true positive rate), or total number of correctly classified observations.
Extended Capabilities
Tall Arrays
Calculate with arrays that have more rows than fit in memory.
This function fully supports tall arrays. For more information, see Tall Arrays.
Version History
Introduced in R2008b
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.

Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list:
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)