Kmedoids
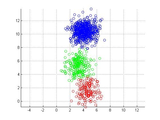
This is a fully vectorized version kmedoids clustering methods (http://en.wikipedia.org/wiki/K-medoids). It is usually more robust than kmeans algorithm. Please try following code for a demo:
close all; clear;
d = 2;
k = 3;
n = 500;
[X,label] = kmeansRnd(d,k,n);
y = kmedoids(X,k);
plotClass(X,label);
figure;
plotClass(X,y);
Input data are assumed COLUMN vectors!
You can only visualize 2d data!
This function is now a part of the PRML toolbox (http://www.mathworks.com/matlabcentral/fileexchange/55826-pattern-recognition-and-machine-learning-toolbox)
Cite As
Mo Chen (2024). Kmedoids (https://www.mathworks.com/matlabcentral/fileexchange/28898-kmedoids), MATLAB Central File Exchange. Retrieved .
MATLAB Release Compatibility
Platform Compatibility
Windows macOS LinuxCategories
Tags
Acknowledgements
Inspired by: Pattern Recognition and Machine Learning Toolbox
Inspired: Parallel Coordinate Plots GUI toolbox
Community Treasure Hunt
Find the treasures in MATLAB Central and discover how the community can help you!
Start Hunting!Discover Live Editor
Create scripts with code, output, and formatted text in a single executable document.