mslowess
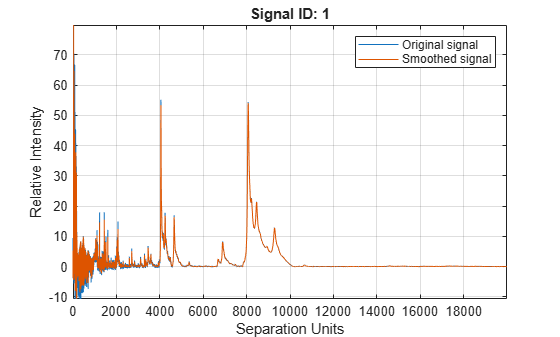
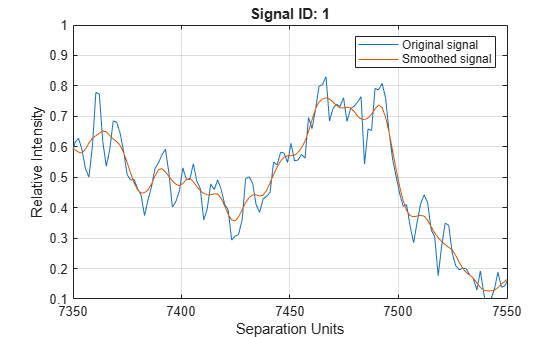
Smooth signal with peaks using nonparametric method
Syntax
Description
mslowess(
smooths raw noisy signal data, X,Intensities)Intensities, using a locally weighted
linear regression (Lowess) method with a default span of 10 samples and
plots the result.
Use mssgolay with data from any separation technique that produces
signal data, such as spectroscopy, NMR, electrophoresis, chromatography, or mass
spectrometry.
mslowess assumes the input vector, X, may not
have uniformly spaced separation units. Therefore, the sliding window for smoothing is
centered using the closest samples in terms of the X value and not in
terms of the X index.
When the input vector, X, does not have repeated values or
NaN values, the algorithm is approximately twice as fast.
Yout = mslowess(X,Intensities)Yout. This syntax does not plot the
data.
Yout = mslowess(X,Intensities,Name=Value)
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Version History
Introduced before R2006a
See Also
mspalign | msbackadj | msdotplot | msalign | msheatmap | msnorm | mspeaks | msresample | msppresample | mssgolay | msviewer
Topics
- Mass Spectrometry and Bioanalytics
- Preprocessing Raw Mass Spectrometry Data
- Visualizing and Preprocessing Hyphenated Mass Spectrometry Data Sets for Metabolite and Protein/Peptide Profiling
- Differential Analysis of Complex Protein and Metabolite Mixtures Using Liquid Chromatography/Mass Spectrometry (LC/MS)