Construct a Simple Model
This example shows you how to construct a simple model with two species (A and B) and a reaction. The reaction is A -> B, which follows mass action kinetics with the forward rate parameter k. Hence the rate of change is .
Create a SimBiology model named simpleModel.
m1 = sbiomodel('simpleModel');Add a reaction that involves two species A and B, where A is converted to B.
r1 = addreaction(m1,'A -> B');SimBiology automatically add species A and B to the model.
m1.species
ans = SimBiology Species Array Index: Compartment: Name: Value: Units: 1 unnamed A 0 2 unnamed B 0
Set the initial amount of the first species (A) to 10.
m1.species(1).InitialAmount = 10;
Define the kinetic law of the reaction to follow mass action kinetics. You can achieve this by adding a kinetic law object to the reaction r1.
kineticLaw = addkineticlaw(r1,'MassAction');Add a rate constant parameter to the mass action kinetic law. You must set the ParameterVariableNames property of the kinetic law object to the name of the parameter 'k' so that the reaction rate can be determined.
p1 = addparameter(kineticLaw,'k',0.5); kineticLaw.ParameterVariableNames = 'k';
Simulate the model.
sd = sbiosimulate(m1);
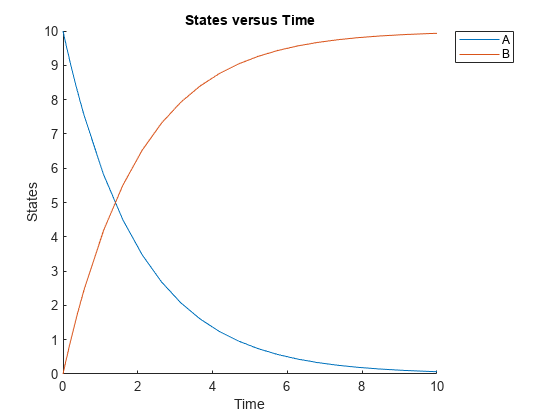
Plot the simulation results.
sbioplot(sd);
See Also
sbiomodel | addreaction (model) | addparameter (model, kineticlaw) | addkineticlaw (reaction) | sbiosimulate