sbiodose
Construct dose object
Syntax
Description
dose = sbiodose(DoseName)RepeatDose object and sets
its Name property to DoseName.
dose = sbiodose(DoseName,DoseType)RepeatDose object or ScheduleDose object based on DoseType.
Examples
This example shows how to set up a dosing regimen that follows the first-order absorption kinetics.
Background
Suppose you have a one-compartment model with a species named drug that represents the total amount of drug in the body. The drug is added to the body via the first-order dosing represented by the reaction dose -> drug, with the absorption rate constant ka. It is removed from the body via the first-order elimination represented by the reaction drug -> null, with the elimination rate constant ke. This example shows how to set up such a one-compartment model, the first-order absorption and elimination.
Create a One-Compartment Model
Create a SimBiology model named onecomp.
m1 = sbiomodel('onecomp');Define the drug elimination by adding a reaction drug -> null to the model. The drug species represents the total amount of drug in the compartment.
r1 = addreaction(m1,'drug -> null');Note that a compartment and the species drug are automatically created, and drug is added to the compartment. The null species is a reserved species that acts as a sink in this reaction.
Add a mass action kinetic law to the reaction. This kinetic law defines the drug elimination to follow the first-order kinetics.
k1 = addkineticlaw(r1,'MassAction');Define the elimination rate parameter ke and add it to the kinetic law.
p1 = addparameter(k1,'ke','Value',1.0,'ValueUnits','1/hour');
Specify the rate parameter ke as the forward rate parameter of the reaction by setting the ParameterVariableNames property of kinetic law object k2. This allows SimBiology to determine the reaction rate for drug -> null reaction.
k1.ParameterVariableNames = 'ke';Set Up the First-Order Dosing
Add a reaction that represents the drug absorption using the second species dose. It represents an intermediate species that will be dosed directly and is required to set up the first-order absorption kinetics.
r2 = addreaction(m1,'dose -> drug');Add a mass action kinetic law to the reaction. This kinetic law defines the drug absorption to follow the first-order kinetics.
k2 = addkineticlaw(r2,'MassAction');Define the absorption rate parameter ka and add it to the kinetic law.
p2 = addparameter(k2,'ka','Value',0.1,'ValueUnits','1/hour');
Specify the rate parameter ka as the forward rate parameter of the reaction by setting the ParameterVariableNames property of kinetic law object k1. This allows SimBiology to determine the reaction rate for dose -> drug reaction.
k2.ParameterVariableNames = 'ka';Suppose you want to increase the drug concentration in the system by administering a series of doses: 250 mg three times a day (t.i.d) for two days. Specify the amount of the dose (Amount), the time interval between each dose (Interval), and the total number of doses (RepeatCount). You also need to set the Active property of the dose object to true so that the dose will be applied to the model during simulation. RepeatCount was set to 5, instead of 6 since it represents the number of doses after the first dose at the default dose start time (d1.StartTime = 0).
d1 = sbiodose('d1','repeat'); d1.Amount = 250; d1.AmountUnits = 'milligram'; d1.Interval = 8; d1.TimeUnits = 'hour'; d1.RepeatCount = 5; d1.Active = true;
Specify the target species of the dose object. The target must be the dose species, not the drug species, so that the drug absorption follows the first-order kinetics.
d1.TargetName = 'dose';Simulate the Model
Change the simulation stop time to 48 hours to match the dosing schedule. Set the unit of compartment to liter.
cs = getconfigset(m1); cs.StopTime = 48; cs.TimeUnits = 'hour'; c1 = m1.Compartments; c1.Units = "liter";
In addition, do not log the dose species data as you are mainly interested in monitoring the drug species which is the drug concentration in the system. This makes visualizing the species in a plot more convenient. To accomplish this, set the StatesToLog property to include the species drug only.
cs.RuntimeOptions.StatesToLog = {'drug'};Simulate the model using the dosing schedule defined by the |d1| dose object.
[t,sd,species] = sbiosimulate(m1,d1);
Plot Results
Plot the concentration versus the time profile of the drug in the compartment.
plot(t,sd); legend(species,'Location','NorthWest'); xlabel('Hours'); ylabel('Drug Concentration');
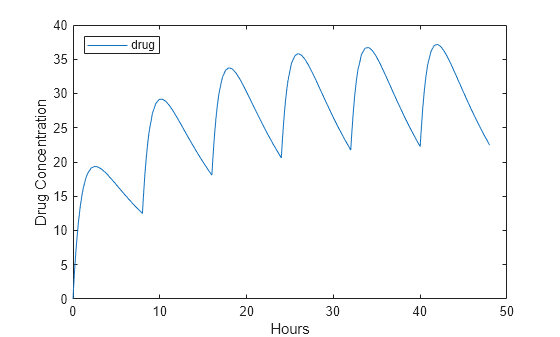
This example shows how to add a series of bolus doses to one-compartment model.
Background
Suppose you have a one-compartment model with a species named drug that represents the total amount of drug in the body. The drug is removed from the body via the first-order elimination represented by the reaction drug -> null, with the elimination rate constant ke. In other words, the drug concentration versus the time profile follows the monoexponential decline , where is the drug concentration at time t, is the initial concentration, and ke is the elimination rate constant. This example shows how to set up such a one-compartment model and administer a series of bolus doses, namely 250 mg three times a day (tid) for two days.
Create a One-Compartment Model
First create a SimBiology model named onecomp.
m1 = sbiomodel('onecomp');Define the elimination of the drug from the system by adding a reaction drug -> null to the model.
r1 = addreaction(m1,'drug -> null');The species drug is automatically created and the reaction is added to the compartment. The null species is a reserved species that acts as a sink in this reaction.
Add a mass action kinetic law to the reaction. This kinetic law defines the drug elimination to follow the first-order kinetics.
k1 = addkineticlaw(r1,'MassAction');Define the elimination rate parameter ke and add it to the kinetic law.
p1 = addparameter(k1,'ke','Value',1.0,'ValueUnits','1/hour');
Specify the rate parameter ke as the forward rate parameter of the reaction by setting the ParameterVariableNames property of kinetic law object k1. This allows SimBiology to determine the reaction rate for drug -> null reaction.
k1.ParameterVariableNames = 'ke';Set Up a Series of Bolus Doses
Suppose you want to increase the drug concentration in the system by administering a series of bolus doses: 250 mg three times a day (tid) for two days. Create a repeat dose object. Specify the amount of the dose (Amount), the dose target, the time interval between each dose (Interval), and the total number of doses (RepeatCount). You also need to set the Active property of the dose object to true so that the dose is applied to the model during simulation.
d1 = sbiodose('d1','repeat'); d1.Amount = 250; d1.AmountUnits = 'milligram'; d1.TargetName = 'drug'; d1.Interval = 8; d1.TimeUnits = 'hour'; d1.RepeatCount = 5; d1.Active = true;
RepeatCount was set to 5, instead of 6 since it represents the number of doses after the first dose at the default dose start time (d1.StartTime = 0).
Simulate the Model
Change the simulation stop time to 48 hours to match the dosing schedule defined by the d1 dose object. Set the compartment unit to liter.
cs = getconfigset(m1); cs.StopTime = 48; cs.TimeUnits = 'hour'; c1 = m1.Compartments; c1.Units = "liter"; [t,sd,species] = sbiosimulate(m1,d1);
Plot Results
Plot the concentration versus the time profile of the drug in the system.
plot(t,sd); legend(species); xlabel('Hours'); ylabel('Drug Concentration');
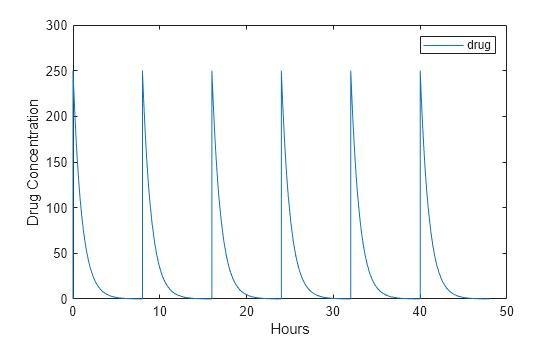
This example shows how to set up a dosing regimen that follows the zero-order absorption kinetics.
Background
Suppose you have a one-compartment model with a species named drug that represents the total amount of drug in the body. The drug is removed from the body via the first-order elimination represented by the reaction drug -> null, with the elimination rate constant ke. In other words, the drug concentration versus the time profile follows the monoexponential decline , where is the drug concentration at time t, is the initial concentration, and ke is the elimination rate constant. This example shows how to set up such a one-compartment model and increase the drug concentration in the compartment via the zero-order absorption that takes 25 hours to administer the total dose amount of 250 mg.
Create a One-Compartment Model
Create a SimBiology model named onecomp.
m1 = sbiomodel('onecomp');Define the elimination of the drug from the system by adding a reaction drug -> null to the model.
r1 = addreaction(m1,'drug -> null');The species drug is automatically created and added to the compartment. The null species is a reserved species that acts as a sink in this reaction.
Add a mass action kinetic law to the reaction. This kinetic law defines the drug elimination to follow the first-order kinetics.
k1 = addkineticlaw(r1,'MassAction');Define the elimination rate parameter ke and add it to the kinetic law.
p1 = addparameter(k1,'ke','Value',1.0,'ValueUnits','1/hour');
Specify the rate parameter ke as the forward rate parameter of the reaction by setting the ParameterVariableNames property of kinetic law object k1. This allows SimBiology to determine the reaction rate for drug -> null reaction.
k1.ParameterVariableNames = 'ke';Set Up Zero-Order Dosing
To set up zero-order dosing, first create a zero-order duration parameter p2 that represents the time it takes to administer a dose. Next, specify the amount of the dose (Amount), the dose target (TargetName), and the name of the zero-order duration parameter (DurationParameterName). You also need to set the Active property of the dose object to true so that the dose is applied to the model during simulation.
p2 = addparameter(m1,'duration','Value',25,'ValueUnits','hour'); d1 = sbiodose('d1'); d1.Amount = 250; d1.AmountUnits = 'milligram'; d1.TargetName = 'drug'; d1.DurationParameterName = 'duration'; %Name of the duration parameter |p2| d1.Active = true;
Simulate the Model
Change the simulation stop time to 48 hours to see the complete time profile. Apply the dosing schedule defined by d1 to the model during simulation.
cs = getconfigset(m1); cs.StopTime = 48; cs.TimeUnits = 'hour'; c1 = m1.Compartments; c1.Units = "liter"; [t,sd,species] = sbiosimulate(m1,d1);
Plot Results
Plot the concentration versus the time profile of the drug in the compartment.
plot(t,sd); legend(species); xlabel('Hours'); ylabel('Drug Concentration');
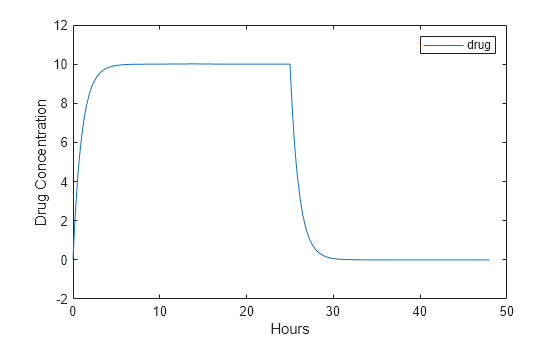
This example shows how to add a constant-rate infusion dose to one-compartment model.
Background
Suppose you have a one-compartment model with a species named drug that represents the total amount of drug in the body. The drug is removed from the body via the first-order elimination represented by the reaction drug -> null, with the elimination rate constant ke. In other words, the drug concentration versus the time profile follows the monoexponential decline , where is the drug concentration at time t, is the initial concentration, and ke is the elimination rate constant. This example shows how to set up such a one-compartment model and add an infusion dose at a constant rate of 10 mg/hour for the total dose amount of 250 mg.
Create a One-Compartment Model
Create a SimBiology model named onecomp.
m1 = sbiomodel('onecomp');Define the elimination of the drug from the system by adding a reaction drug -> null to the model.
r1 = addreaction(m1,'drug -> null');The species drug is automatically created and added to the compartment. The null species is a reserved species that acts as a sink in this reaction.
Add a mass action kinetic law to the reaction. This kinetic law defines the drug elimination to follow the first-order kinetics.
k1 = addkineticlaw(r1,'MassAction');Define the elimination rate parameter ke and add it to the kinetic law.
p1 = addparameter(k1,'ke','Value',1.0,'ValueUnits','1/hour');
Specify the rate parameter ke as the forward rate parameter of the reaction by setting the ParameterVariableNames property of kinetic law object k1. This allows SimBiology to determine the reaction rate for drug -> null reaction.
k1.ParameterVariableNames = 'ke';Set Up an Infusion Dose
Specify the amount of the dose (Amount), the dose target (TargetName), and the infusion rate (Rate). You also need to set the Active property of the dose object to true so that the dose is applied to the model during simulation.
d1 = sbiodose('d1'); d1.Amount = 250; d1.TargetName = 'drug'; d1.Rate = 10; d1.RateUnits = 'milligram/hour'; d1.Active = true;
Simulate the Model
Change the simulation stop time to 48 hours to see the complete time course. Apply the dosing schedule defined by d1 to the model during simulation.
cs = getconfigset(m1); cs.StopTime = 48; cs.TimeUnits = 'hour'; c1 = m1.Compartments; c1.Units = "liter"; [t,sd,species] = sbiosimulate(m1,d1);
Plot Results
Plot the concentration versus the time profile of the drug in the system.
plot(t,sd); legend(species); xlabel('Hours'); ylabel('Drug Concentration');
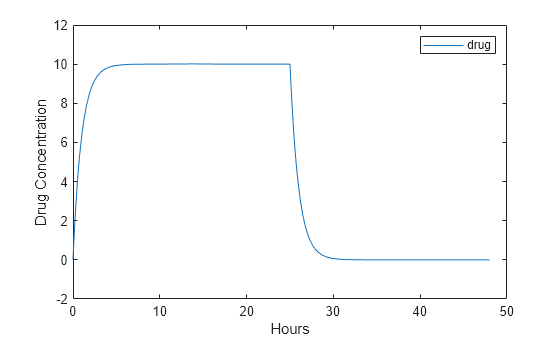
Input Arguments
Name of the dose object, specified as a character vector or string.
Example: '250mg_tid'
Data Types: char
Type of the dose object, specified as 'schedule' for
a ScheduleDose object and 'repeat' for
a RepeatDose object.
Example: 'schedule'
Data Types: char
Output Arguments
Dose object, returned as a RepeatDose
object or ScheduleDose object.
Version History
Introduced in R2010a
See Also
adddose | getdose | removedose | copyobj | get | set | ScheduleDose
object | RepeatDose object
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)