predict
Predict labels using discriminant analysis classifier
Description
[ also returns:label,score,cost]
= predict(Mdl,X)
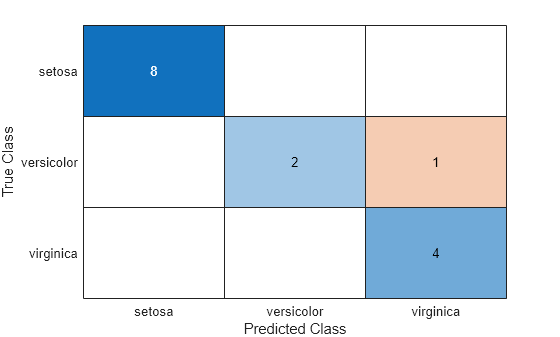
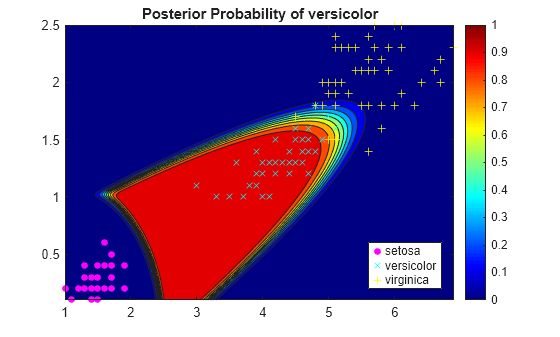
A matrix of classification scores (
score) indicating the likelihood that a label comes from a particular class. For discriminant analysis, scores are posterior probabilities.A matrix of expected classification cost (
cost). For each observation inX, the predicted class label corresponds to the minimum expected classification cost among all classes.
Examples
Input Arguments
Output Arguments
More About
Alternative Functionality
Simulink Block
To integrate the prediction of a discriminant analysis classification model into
Simulink®, you can use the ClassificationDiscriminant
Predict block in the Statistics and Machine Learning Toolbox™ library or a MATLAB® Function block with the predict function. For examples,
see Predict Class Labels Using ClassificationDiscriminant Predict Block and Predict Class Labels Using MATLAB Function Block.
When deciding which approach to use, consider the following:
If you use the Statistics and Machine Learning Toolbox library block, you can use the Fixed-Point Tool (Fixed-Point Designer) to convert a floating-point model to fixed point.
Support for variable-size arrays must be enabled for a MATLAB Function block with the
predictfunction.If you use a MATLAB Function block, you can use MATLAB functions for preprocessing or post-processing before or after predictions in the same MATLAB Function block.
Extended Capabilities
Version History
Introduced in R2011b