glmfit
Fit generalized linear regression model
Syntax
Description
b = glmfit(X,y,distr,Name,Value)'Constant','off' to omit the constant term from the model.
Examples
Fit Generalized Linear Model with Probit Link
Fit a generalized linear regression model, and compute predicted (estimated) values for the predictor data using the fitted model.
Create a sample data set.
x = [2100 2300 2500 2700 2900 3100 ...
3300 3500 3700 3900 4100 4300]';
n = [48 42 31 34 31 21 23 23 21 16 17 21]';
y = [1 2 0 3 8 8 14 17 19 15 17 21]';x contains the predictor variable values. Each y value is the number of successes in the corresponding number of trials in n.
Fit a probit regression model for y on x.
b = glmfit(x,[y n],'binomial','Link','probit');
Compute the estimated number of successes.
yfit = glmval(b,x,'probit','Size',n);
Plot the observed success percent and estimated success percent versus the x values.
plot(x,y./n,'o',x,yfit./n,'-')
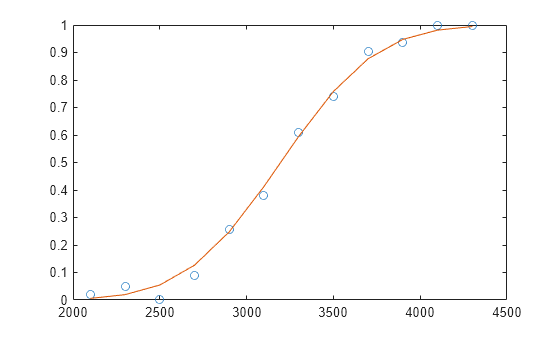
Fit Generalized Linear Model Using Custom Link Function
Define a custom link function and use it to fit a generalized linear regression model.
Load the sample data.
load fisheririsThe column vector species contains iris flowers of three different species: setosa, versicolor, and virginica. The matrix meas contains four types of measurements for the flowers, the length and width of sepals and petals in centimeters.
Define the predictor variables and response variable.
X = meas(51:end,:);
y = strcmp('versicolor',species(51:end));Define a custom link function for a logit link function. Create three function handles that define the link function, the derivative of the link function, and the inverse link function. Store them in a cell array.
link = @(mu) log(mu./(1-mu));
derlink = @(mu) 1./(mu.*(1-mu));
invlink = @(resp) 1./(1+exp(-resp));
F = {link,derlink,invlink};Fit a logistic regression model using glmfit with the custom link function.
b = glmfit(X,y,'binomial','link',F)
b = 5×1
42.6378
2.4652
6.6809
-9.4294
-18.2861
Fit a generalized linear model by using the built-in logit link function, and compare the results.
b = glmfit(X,y,'binomial','link','logit')
b = 5×1
42.6378
2.4652
6.6809
-9.4294
-18.2861
Perform Deviance Test
Fit a generalized linear regression model that contains an intercept and linear term for each predictor. Perform a deviance test that determines whether the model fits significantly better than a constant model.
Generate sample data using Poisson random numbers with two underlying predictors X(:,1) and X(:,2).
rng('default') % For reproducibility rndvars = randn(100,2); X = [2 + rndvars(:,1),rndvars(:,2)]; mu = exp(1 + X*[1;2]); y = poissrnd(mu);
Fit a generalized linear regression model that contains an intercept and linear term for each predictor.
[b,dev] = glmfit(X,y,'poisson');The second output argument dev is a Deviance of the fit.
Fit a generalized linear regression model that contains only an intercept. Specify the predictor variable as a column of 1s, and specify 'Constant' as 'off' so that glmfit does not include a constant term in the model.
[~,dev_noconstant] = glmfit(ones(100,1),y,'poisson','Constant','off');
Compute the difference between dev_constant and dev.
D = dev_noconstant - dev
D = 2.9533e+05
D has a chi-square distribution with 2 degrees of freedom. The degrees of freedom equal the difference in the number of estimated parameters in the model corresponding to dev and the number of estimated parameters in the constant model. Find the p-value for a deviance test.
p = 1 - chi2cdf(D,2)
p = 0
The small p-value indicates that the model differs significantly from a constant.
Alternatively, you can create a generalized linear regression model of Poisson data by using the fitglm function. The model display includes the statistic (Chi^2-statistic vs. constant model) and p-value.
mdl = fitglm(X,y,'y ~ x1 + x2','Distribution','poisson')
mdl =
Generalized linear regression model:
log(y) ~ 1 + x1 + x2
Distribution = Poisson
Estimated Coefficients:
Estimate SE tStat pValue
________ _________ ______ ______
(Intercept) 1.0405 0.022122 47.034 0
x1 0.9968 0.003362 296.49 0
x2 1.987 0.0063433 313.24 0
100 observations, 97 error degrees of freedom
Dispersion: 1
Chi^2-statistic vs. constant model: 2.95e+05, p-value = 0
You can also use the devianceTest function with the fitted model object.
devianceTest(mdl)
ans=2×4 table
Deviance DFE chi2Stat pValue
__________ ___ __________ ______
log(y) ~ 1 2.9544e+05 99
log(y) ~ 1 + x1 + x2 107.4 97 2.9533e+05 0
Input Arguments
X — Predictor variables
numeric matrix
Predictor variables, specified as an n-by-p
numeric matrix, where n is the number of observations and
p is the number of predictor variables. Each column of
X represents one variable, and each row represents one
observation.
By default, glmfit includes a constant term in the model. Do
not add a column of 1s directly to X. You can change the default
behavior of glmfit by specifying the
'Constant' name-value argument.
Data Types: single | double
y — Response variable
vector | matrix
Response variable, specified as a vector or matrix.
If
distris not'binomial', thenymust be an n-by-1 vector, where n is the number of observations. Each entry inyis the response for the corresponding row ofX. The data type must be single or double.If
distris'binomial', thenyis an n-by-1 vector indicating success or failure at each observation, or an n-by-2 matrix whose first column indicates the number of successes for each observation and second column indicates the number of trials for each observation.
Data Types: single | double | logical | categorical
distr — Distribution of response variable
'normal' (default) | 'binomial' | 'poisson' | 'gamma' | 'inverse gaussian'
Distribution of the response variable, specified as one of the values in this table.
| Value | Description |
|---|---|
'normal' | Normal distribution (default) |
'binomial' | Binomial distribution |
'poisson' | Poisson distribution |
'gamma' | Gamma distribution |
'inverse gaussian' | Inverse Gaussian distribution |
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: b = glmfit(X,y,'normal','link','probit') specifies that the
distribution of the response is normal and instructs glmfit to use the
probit link function.
B0 — Initial values for coefficient estimates
numeric vector
Initial values for the coefficient estimates, specified as a numeric vector. The default values are initial fitted values derived from the input data.
Data Types: single | double
Constant — Indicator for constant term
'on' (default) | 'off'
Indicator for the constant term (intercept) in the fit, specified as either
'on' to include the constant term or 'off' to
remove it from the model.
'on'(default) —glmfitincludes a constant term in the model and returns a (p + 1)-by-1 vector of coefficient estimatesb, where p is the number of predictors inX. The coefficient of the constant term is the first element ofb.'off'—glmfitomits the constant term and returns a p-by-1 vector of coefficient estimatesb.
Example: 'Constant','off'
EstDisp — Indicator to compute dispersion parameter
'off' for 'binomial' and
'poisson' distributions (default) | 'on'
Indicator to compute a dispersion parameter for 'binomial' and
'poisson' distributions, specified as 'on' or
'off'.
| Value | Description |
|---|---|
'on' | Estimate a dispersion parameter when computing standard errors. The estimated dispersion parameter value is the sum of squared Pearson residuals divided by the degrees of freedom for error (DFE). |
'off' | Use the theoretical value of 1 when computing standard errors (default). |
The fitting function always estimates the dispersion for other distributions.
Example: 'EstDisp','on'
LikelihoodPenalty — Penalty for likelihood estimate
"none" (default) | "jeffreys-prior"
Penalty for the likelihood estimate, specified as "none" or
"jeffreys-prior".
"none"—glmfitdoes not apply a penalty to the likelihood estimate."jeffreys-prior"—glmfituses Jeffreys prior to penalize the likelihood estimate.
For logistic models, setting LikelihoodPenalty to
"jeffreys-prior" is called Firth's
regression. To reduce the coefficient estimate bias when you have a small
number of samples, or when you are performing binomial (logistic) regression on a
separable data set, set LikelihoodPenalty to
"jeffreys-prior".
Example: LikelihoodPenalty="jeffreys-prior"
Data Types: char | string
Link — Link function
canonical link function (default) | scalar value | structure or cell array of custom link function
Link function to use in place of the canonical link function, specified as one of the built-in link functions in the following table or a custom link function.
| Link Function Name | Link Function | Mean (Inverse) Function |
|---|---|---|
'identity' (default for 'normal'
distribution) | f(μ) = μ | μ = Xb |
'log' (default for 'poisson'
distribution) | f(μ) = log(μ) | μ = exp(Xb) |
'logit' (default for 'binomial'
distribution) | f(μ) = log(μ/(1 – μ)) | μ = exp(Xb) / (1 + exp(Xb)) |
'probit' | f(μ) = Φ–1(μ), where Φ is the cumulative distribution function of the standard normal distribution | μ = Φ(Xb) |
| f(μ) = log(–log(μ)) | μ = exp(–exp(Xb)) |
'comploglog' | f(μ) = log(–log(1 – μ)) | μ = 1 – exp(–exp(Xb)) |
'reciprocal' (default for 'gamma'
distribution) | f(μ) = 1/μ | μ = 1/(Xb) |
p (a number, default for the 'inverse
gaussian' distribution with p = –2) | f(μ) = μp | μ = Xb1/p |
The default 'Link' value is the canonical link
function, which depends on the distribution of the response variable specified by the
distr argument.
You can specify a custom link function using a structure or cell array.
Structure with three fields. Each field of the structure (for example,
S) holds a function handle that accepts a vector of inputs and returns a vector of the same size:S.Link— Link function, f(μ) =S.Link(μ)S.Derivative— Derivative of the link functionS.Inverse— Inverse link function, μ =S.Inverse(Xb)
Cell array of the form
{FL FD FI}that defines the link function (FL(mu)), the derivative of the link function (FD = dFL(mu)/dmu), and the inverse link function (FI = FL^(–1)). Each entry holds a function handle that accepts a vector of inputs and returns a vector of the same size.
The link function defines the relationship f(μ) = X*b between the mean response μ and the linear combination of predictors X*b.
Example: 'Link','probit'
Data Types: single | double | char | string | struct | cell
Offset — Offset variable
[ ] (default) | numeric vector
Offset variable in the fit, specified as a numeric vector with the same length as
the response y.
glmfit uses Offset as an additional
predictor with a coefficient value fixed at 1. In other words, the formula for fitting is
f(μ) = Offset +
X*b,
where f is the link function,
μ is the mean response, and
X*b is the linear combination of predictors
X. The Offset predictor has coefficient
1.
For example, consider a Poisson regression model. Suppose, for theoretical
reasons, the number of counts is to be proportional to a predictor
A. By using the log link function and specifying
log(A) as an offset, you can force the model to satisfy this
theoretical constraint.
Data Types: single | double
Options — Optimization options
statset('glmfit') (default) | structure
glmfit')Optimization options, specified as a structure. This argument determines the control
parameters for the iterative algorithm that glmfit
uses.
Create the 'Options' value by using the function statset or by creating a structure array containing the fields and values described in this table.
| Field Name | Value | Default Value |
|---|---|---|
Display | Amount of information displayed by the algorithm
| 'off' |
MaxIter | Maximum number of iterations allowed, specified as a positive integer | 100 |
TolX | Termination tolerance for the parameters, specified as a positive scalar | 1e-6 |
You can also enter statset(' in the
Command Window to see the names and default values of the fields that
glmfit')glmfit accepts in the 'Options'
name-value argument.
Example: 'Options',statset('Display','final','MaxIter',1000) specifies to display the final information of the iterative algorithm results, and change the maximum number of iterations allowed to 1000.
Data Types: struct
Weights — Observation weights
ones(n,1) (default) | n-by-1 vector of nonnegative scalar values
Observation weights, specified as an n-by-1 vector of nonnegative scalar values, where n is the number of observations.
Data Types: single | double
Output Arguments
b — Coefficient estimates
numeric vector
Coefficient estimates, returned as a numeric vector.
If
'Constant'is'on'(default), thenglmfitincludes a constant term in the model and returns a (p + 1)-by-1 vector of coefficient estimatesb, where p is the number of predictors inX. The coefficient of the constant term is the first element ofb.If
'Constant'is'off', thenglmfitomits the constant term and returns a p-by-1 vector of coefficient estimatesb.
dev — Deviance of fit
numeric value
Deviance of the fit, returned as a numeric value. The deviance is useful for comparing two models when one model is a special case of the other model. The difference between the deviance of the two models has a chi-square distribution with degrees of freedom equal to the difference in the number of estimated parameters between the two models.
For more information, see Deviance.
stats — Model statistics
structure
Model statistics, returned as a structure with the following fields:
beta— Coefficient estimatesbdfe— Degrees of freedom for errorsfit— Estimated dispersion parameters— Theoretical or estimated dispersion parameterestdisp— 0 when'EstDisp'is'off'and 1 when'EstDisp'is'on'covb— Estimated covariance matrix forbse— Vector of standard errors of the coefficient estimatesbcoeffcorr— Correlation matrix forbt— t statistics forbp— p-values forbresid— Vector of residualsresidp— Vector of Pearson residualsresidd— Vector of deviance residualsresida— Vector of Anscombe residuals
If you estimate a dispersion parameter for the binomial or Poisson distribution,
then stats.s is equal to stats.sfit. Also, the
elements of stats.se differ by the factor stats.s
from their theoretical values.
More About
Deviance
Deviance is a generalization of the residual sum of squares. It measures the goodness of fit compared to a saturated model.
The deviance of a model M1 is twice the difference between the loglikelihood of the model M1 and the saturated model Ms. A saturated model is a model with the maximum number of parameters that you can estimate.
For example, if you have n observations (yi, i = 1, 2, ..., n) with potentially different values for XiTβ, then you can define a saturated model with n parameters. Let L(b,y) denote the maximum value of the likelihood function for a model with the parameters b. Then the deviance of the model M1 is
where b1 and bs contain the estimated parameters for the model M1 and the saturated model, respectively. The deviance has a chi-squared distribution with n – p degrees of freedom, where n is the number of parameters in the saturated model and p is the number of parameters in the model M1.
Assume you have two different generalized linear regression models M1 and M2, and M1 has a subset of the terms in M2. You can assess the fit of the models by comparing their deviances D1 and D2. The difference of the deviances is
Asymptotically, the difference D has a chi-squared distribution with
degrees of freedom v equal to the difference in the number of parameters
estimated in M1 and
M2. You can obtain the
p-value for this test by using 1 —
chi2cdf(D,v).
Typically, you examine D using a model M2 with a constant term and no predictors. Therefore, D has a chi-squared distribution with p – 1 degrees of freedom. If the dispersion is estimated, the difference divided by the estimated dispersion has an F distribution with p – 1 numerator degrees of freedom and n – p denominator degrees of freedom.
Alternative Functionality
glmfit is useful when you simply need the output arguments of the
function or when you want to repeat fitting a model multiple times in a loop. If you need to
investigate a fitted model further, create a generalized linear regression model object GeneralizedLinearModel by using fitglm or stepwiseglm. A
GeneralizedLinearModel object provides more features than
glmfit.
Use the properties of
GeneralizedLinearModelto investigate a fitted model. The object properties include information about the coefficient estimates, summary statistics, fitting method, and input data.Use the object functions of
GeneralizedLinearModelto predict responses and to modify, evaluate, and visualize the generalized linear regression model.You can find the information in the output of
glmfitusing the properties and object functions ofGeneralizedLinearModel.Output of glmfitEquivalent Values in GeneralizedLinearModelbSee the Estimatecolumn of theCoefficientsproperty.devSee the Devianceproperty.statsSee the model display in the Command Window. You can find the statistics in the model properties (
CoefficientCovariance,Coefficients,Dispersion,DispersionEstimated, andResiduals).The dispersion parameter in
stats.sglmfitis the scale factor for the standard errors of coefficients, whereas the dispersion parameter in theDispersionproperty of a generalized linear model is the scale factor for the variance of the response. Therefore,stats.sis the square root of theDispersionvalue.
References
[1] Dobson, A. J. An Introduction to Generalized Linear Models. New York: Chapman & Hall, 1990.
[2] McCullagh, P., and J. A. Nelder. Generalized Linear Models. New York: Chapman & Hall, 1990.
[3] Collett, D. Modeling Binary Data. New York: Chapman & Hall, 2002.
Extended Capabilities
GPU Arrays
Accelerate code by running on a graphics processing unit (GPU) using Parallel Computing Toolbox™.
This function fully supports GPU arrays. For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced before R2006a
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.

Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list:
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)