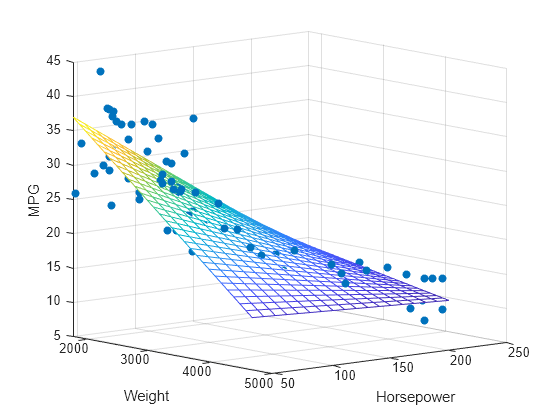
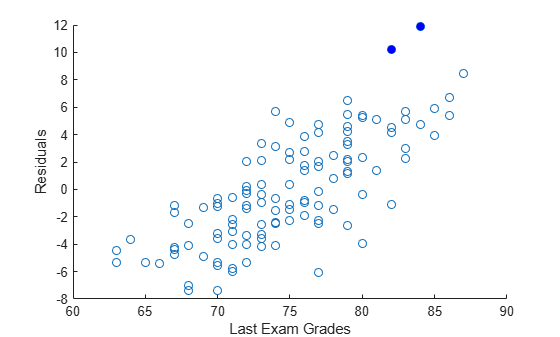
regress
Multiple linear regression
Syntax
Description
Examples
Input Arguments
Output Arguments
Tips
Algorithms
Alternative Functionality
regress is useful when you simply need the output arguments of
the function and when you want to repeat fitting a model multiple times in a loop. If
you need to investigate a fitted regression model further, create a linear regression
model object LinearModel by using fitlm or stepwiselm. A LinearModel
object provides more features than regress.
Use the properties of
LinearModelto investigate a fitted linear regression model. The object properties include information about coefficient estimates, summary statistics, fitting method, and input data.Use the object functions of
LinearModelto predict responses and to modify, evaluate, and visualize the linear regression model.Unlike
regress, thefitlmfunction does not require a column of ones in the input data. A model created byfitlmalways includes an intercept term unless you specify not to include it by using the'Intercept'name-value pair argument.You can find the information in the output of
regressusing the properties and object functions ofLinearModel.Output of regressEquivalent Values in LinearModelbSee the Estimatecolumn of theCoefficientsproperty.bintUse the coefCIfunction.rSee the Rawcolumn of theResidualsproperty.rintNot supported. Instead, use studentized residuals ( Residualsproperty) and observation diagnostics (Diagnosticsproperty) to find outliers.statsSee the model display in the Command Window. You can find the statistics in the model properties ( MSEandRsquared) and by using theanovafunction.
References
[1] Chatterjee, S., and A. S. Hadi. “Influential Observations, High Leverage Points, and Outliers in Linear Regression.” Statistical Science. Vol. 1, 1986, pp. 379–416.
Extended Capabilities
Version History
Introduced before R2006a
See Also
LinearModel | fitlm | stepwiselm | mvregress | rcoplot