mahal
Mahalanobis distance to Gaussian mixture component
Syntax
Description
Examples
Generate random variates that follow a mixture of two bivariate Gaussian distributions by using the mvnrnd function. Fit a Gaussian mixture model (GMM) to the generated data by using the fitgmdist function, and then compute Mahalanobis distances between the generated data and the mixture components of the fitted GMM.
Define the distribution parameters (means and covariances) of two bivariate Gaussian mixture components.
rng('default') % For reproducibility mu1 = [1 2]; % Mean of the 1st component sigma1 = [2 0; 0 .5]; % Covariance of the 1st component mu2 = [-3 -5]; % Mean of the 2nd component sigma2 = [1 0; 0 1]; % Covariance of the 2nd component
Generate an equal number of random variates from each component, and combine the two sets of random variates.
r1 = mvnrnd(mu1,sigma1,1000); r2 = mvnrnd(mu2,sigma2,1000); X = [r1; r2];
The combined data set X contains random variates following a mixture of two bivariate Gaussian distributions.
Fit a two-component GMM to X.
gm = fitgmdist(X,2)
gm = Gaussian mixture distribution with 2 components in 2 dimensions Component 1: Mixing proportion: 0.500000 Mean: -2.9617 -4.9727 Component 2: Mixing proportion: 0.500000 Mean: 0.9539 2.0261
fitgmdist fits a GMM to X using two mixture components. The means of Component 1 and Component 2 are [-2.9617,-4.9727] and [0.9539,2.0261], which are close to mu2 and mu1, respectively.
Compute the Mahalanobis distance of each point in X to each component of gm.
d2 = mahal(gm,X);
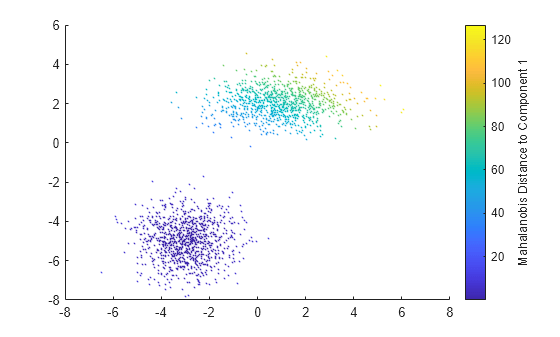
Plot X by using scatter and use marker color to visualize the Mahalanobis distance to Component 1.
scatter(X(:,1),X(:,2),10,d2(:,1),'.') % Scatter plot with points of size 10 c = colorbar; ylabel(c,'Mahalanobis Distance to Component 1')
Input Arguments
Gaussian mixture distribution, also called Gaussian mixture model (GMM), specified as a gmdistribution object.
You can create a gmdistribution object using gmdistribution or fitgmdist. Use the gmdistribution function to create a
gmdistribution object by specifying the distribution parameters.
Use the fitgmdist function to fit a gmdistribution
model to data given a fixed number of components.
Data, specified as an n-by-m numeric matrix, where n is the number of observations and m is the number of variables in each observation.
If a row of X contains NaNs, then
mahal excludes the row from the computation.
The corresponding value in d2 is
NaN.
Data Types: single | double
Output Arguments
Squared Mahalanobis distance of each observation in X to each Gaussian
mixture component in gm, returned as an
n-by-k numeric matrix, where
n is the number of observations in X and
k is the number of mixture components in
gm.
d2(i,j) is the squared distance of observation i to the
jth Gaussian mixture component.
More About
The Mahalanobis distance is a measure between a sample point and a distribution.
The Mahalanobis distance from a vector x to a distribution with mean μ and covariance Σ is
This distance represents how far x is from the mean in number of standard deviations.
mahal returns the squared Mahalanobis distance d2 from an observation in X to a mixture
component in gm.
Version History
Introduced in R2007b
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)