tsne
t-Distributed Stochastic Neighbor Embedding
Description
Y = tsne(X,Name,Value)
Examples
The Fisher iris data set has four-dimensional measurements of irises, and corresponding classification into species. Visualize this data by reducing the dimension using tsne.
load fisheriris rng default % for reproducibility Y = tsne(meas); gscatter(Y(:,1),Y(:,2),species)
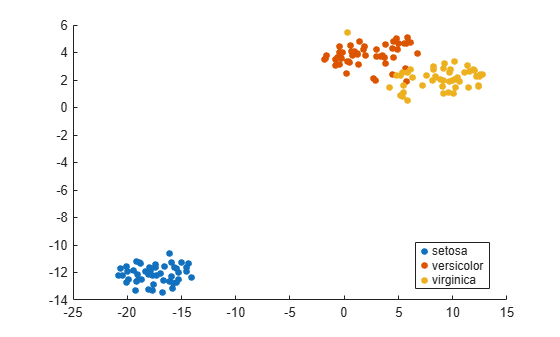
Use various distance metrics to try to obtain a better separation between species in the Fisher iris data.
load fisheriris rng('default') % for reproducibility Y = tsne(meas,'Algorithm','exact','Distance','mahalanobis'); subplot(2,2,1) gscatter(Y(:,1),Y(:,2),species) title('Mahalanobis') rng('default') % for fair comparison Y = tsne(meas,'Algorithm','exact','Distance','cosine'); subplot(2,2,2) gscatter(Y(:,1),Y(:,2),species) title('Cosine') rng('default') % for fair comparison Y = tsne(meas,'Algorithm','exact','Distance','chebychev'); subplot(2,2,3) gscatter(Y(:,1),Y(:,2),species) title('Chebychev') rng('default') % for fair comparison Y = tsne(meas,'Algorithm','exact','Distance','euclidean'); subplot(2,2,4) gscatter(Y(:,1),Y(:,2),species) title('Euclidean')
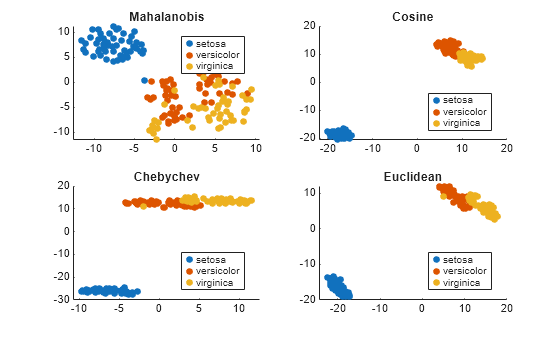
In this case, the cosine, Chebychev, and Euclidean distance metrics give reasonably good separation of clusters. But the Mahalanobis distance metric does not give a good separation.
tsne removes input data rows that contain any NaN entries. Therefore, you must remove any such rows from your classification data before plotting.
For example, change a few random entries in the Fisher iris data to NaN.
load fisheriris rng default % for reproducibility meas(rand(size(meas)) < 0.05) = NaN;
Embed the four-dimensional data into two dimensions using tsne.
Y = tsne(meas,'Algorithm','exact');
Warning: Rows with NaN missing values in X or 'InitialY' values are removed.
Determine how many rows were eliminated from the embedding.
length(species)-length(Y)
ans = 22
Prepare to plot the result by locating the rows of meas that have no NaN values.
goodrows = not(any(isnan(meas),2));
Plot the results using only the rows of species that correspond to rows of meas with no NaN values.
gscatter(Y(:,1),Y(:,2),species(goodrows))
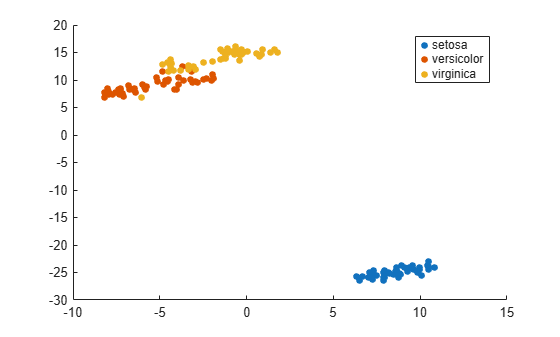
Find both 2-D and 3-D embeddings of the Fisher iris data, and compare the loss for each embedding. It is likely that the loss is lower for a 3-D embedding, because this embedding has more freedom to match the original data.
load fisheriris rng default % for reproducibility [Y,loss] = tsne(meas,'Algorithm','exact'); rng default % for fair comparison [Y2,loss2] = tsne(meas,'Algorithm','exact','NumDimensions',3); fprintf('2-D embedding has loss %g, and 3-D embedding has loss %g.\n',loss,loss2)
2-D embedding has loss 0.1255, and 3-D embedding has loss 0.0980872.
As expected, the 3-D embedding has lower loss.
View the embeddings. Use RGB colors [1 0 0], [0 1 0], and [0 0 1].
For the 3-D plot, convert the species to numeric values using the categorical command, then convert the numeric values to RGB colors using the sparse function as follows. If v is a vector of positive integers 1, 2, or 3, corresponding to the species data, then the command
sparse(1:numel(v),v,ones(size(v)))
is a sparse matrix whose rows are the RGB colors of the species.
gscatter(Y(:,1),Y(:,2),species,eye(3))
title('2-D Embedding')
figure v = double(categorical(species)); c = full(sparse(1:numel(v),v,ones(size(v)),numel(v),3)); scatter3(Y2(:,1),Y2(:,2),Y2(:,3),15,c,'filled') title('3-D Embedding') view(-50,8)
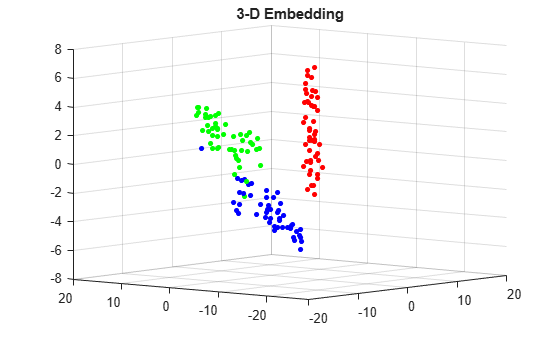
Input Arguments
Data points, specified as an n-by-m matrix,
where each row is one m-dimensional point.
tsne removes rows of X that
contain any NaN values before creating an embedding.
See Plot Results with NaN Input Data.
Data Types: single | double
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: Y =
tsne(X,'Algorithm','Exact','NumPCAComponents',50)
Algorithm Control
tsne algorithm, specified as 'barneshut' or 'exact'.
The 'exact' algorithm optimizes the Kullback-Leibler
divergence of distributions between the original space and the embedded
space. The 'barneshut' algorithm performs an approximate
optimization that is faster and uses less memory when the number of
data rows is large.
Note
For the 'barneshut' algorithm, tsne uses knnsearch to find the nearest neighbors.
Example: 'exact'
Size of the Gram matrix in megabytes, specified as a positive scalar or
"maximal". This argument is valid only when the
Distance name-value argument begins with
fast.
If you set CacheSize to "maximal",
tsne tries to allocate enough memory for an entire
intermediate matrix whose size is M-by-M, where
M is the number of rows of the input data X.
The cache size does not have to be large enough for an entire intermediate matrix, but
must be at least large enough to hold an M-by-1 vector. Otherwise,
tsne uses the standard algorithm for computing Euclidean
distances.
If the value of CacheSize is too large or
"maximal", tsne might try to allocate
a Gram matrix that exceeds the available memory. In this case, the software issues an
error.
Example: CacheSize="maximal"
Data Types: double | char | string
Distance metric, specified as one of the following:
'euclidean'— Euclidean distance.'seuclidean'— Standardized Euclidean distance. Each coordinate difference between the rows inXand the query matrix is scaled by dividing by the corresponding element of the standard deviation computed fromS = std(X,'omitnan').'fasteuclidean'— Euclidean distance computed by using an alternative algorithm that saves time when the number of predictors is at least 10. In some cases, this faster algorithm can reduce accuracy. Algorithms starting with'fast'do not support sparse data. For details, see Algorithms.'fastseuclidean'— Standardized Euclidean distance computed by using an alternative algorithm that saves time when the number of predictors is at least 10. In some cases, this faster algorithm can reduce accuracy. Algorithms starting with'fast'do not support sparse data. For details, see Algorithms.'cityblock'— City block distance.'chebychev'— Chebychev distance, which is the maximum coordinate difference.'minkowski'— Minkowski distance with exponent 2. This distance is the same as the Euclidean distance.'mahalanobis'— Mahalanobis distance, computed using the positive definite covariance matrixcov(X,'omitrows').'cosine'— One minus the cosine of the included angle between observations (treated as vectors).'correlation'— One minus the sample linear correlation between observations (treated as sequences of values).'spearman'— One minus the sample Spearman's rank correlation between observations (treated as sequences of values).'hamming'— Hamming distance, which is the percentage of coordinates that differ.'jaccard'— One minus the Jaccard coefficient, which is the percentage of nonzero coordinates that differ.Custom distance function — A distance function specified using
@(for example,@distfun). For details, see More About.
In all cases, tsne uses squared pairwise
distances to calculate the Gaussian kernel in the joint distribution
of X.
Example: 'mahalanobis'
Data Types: char | string | function_handle
Size of natural clusters in data, specified as a scalar value 1 or
greater.
A large exaggeration makes tsne learn larger
joint probabilities of Y and creates relatively
more space between clusters in Y. tsne uses
exaggeration in the first 99 optimization iterations.
If the value of Kullback-Leibler divergence increases in the early stage of the optimization, try reducing the exaggeration. See Modify t-SNE Settings.
Example: 10
Data Types: single | double
Dimension of the output Y, specified as
a positive integer. Generally, set NumDimensions to 2 or 3.
Example: 3
Data Types: single | double
PCA dimension reduction, specified as a nonnegative integer.
Before tsne embeds the high-dimensional data,
it first reduces the dimensionality of the data to NumPCAComponents using
the pca function. When NumPCAComponents is 0, tsne does
not use PCA.
Example: 50
Data Types: single | double
Effective number of local neighbors of each point, specified as a positive scalar. See t-SNE Algorithm.
Larger perplexity causes tsne to use more
points as nearest neighbors. Use a larger value of Perplexity for
a large dataset. Typical Perplexity values are
from 5 to 50. In the Barnes-Hut
algorithm, tsne uses min(3*Perplexity,N-1) as
the number of nearest neighbors. See Modify t-SNE Settings.
Example: 10
Data Types: single | double
Flag to normalize input data, specified as a numeric or logical 0 (false) or 1 (true). When the value is true, tsne centers and scales each column of X by first subtracting its mean, and then dividing by its standard deviation.
When features in X are on different scales, set Standardize to true. The learning process is based on nearest neighbors, so features with large scales can override the contribution of features with small scales.
Example: Standardize=true
Data Types: logical
Optimization Control
Initial embedded points, specified as an n-by-NumDimensions real
matrix, where n is the number of rows of X.
The tsne optimization algorithm uses these points
as initial values.
Data Types: single | double
Learning rate for optimization process, specified as a positive
scalar. Typically, set values from 100 through 1000.
When LearnRate is too small, tsne can
converge to a poor local minimum. When LearnRate is
too large, the optimization can initially have the Kullback-Leibler
divergence increase rather than decrease. See Modify t-SNE Settings.
Example: 1000
Data Types: single | double
Iterative display frequency, specified as a positive integer.
When the Verbose name-value pair is not 0, tsne returns
iterative display after every NumPrint iterations.
If the Options name-value pair contains a nonempty 'OutputFcn' entry,
then output functions run after every NumPrint iterations.
Example: 20
Data Types: single | double
Optimization options, specified as a structure containing the
fields 'MaxIter', 'OutputFcn',
and 'TolFun'. Create 'Options' using statset or struct.
'MaxIter'— Positive integer specifying the maximum number of optimization iterations. Default:1000.'OutputFcn'— Function handle or cell array of function handles specifying one or more functions to call after everyNumPrintoptimization iterations. For syntax details, see t-SNE Output Function. Default:[].'TolFun'— Stopping criterion for the optimization. The optimization exits when the norm of the gradient of the Kullback-Leibler divergence is less than'TolFun'. Default:1e-10.
Example: options = statset('MaxIter',500)
Data Types: struct
Barnes-Hut tradeoff parameter, specified as a scalar from 0
through 1. Higher values give a faster but less accurate optimization.
Applies only when Algorithm is 'barneshut'.
Example: 0.1
Data Types: single | double
Iterative display, specified as 0, 1,
or 2. When Verbose is not 0, tsne prints
a summary table of the Kullback-Leibler divergence and the norm of
its gradient every NumPrint iterations.
When Verbose is 2, tsne also
prints the variances of Gaussian kernels. tsne uses
these kernels in its computation of the joint probability of X.
If you see a large difference in the scales of the minimum and maximum
variances, you can sometimes get more suitable results by rescaling X.
Example: 2
Data Types: single | double
Output Arguments
Embedded points, returned as an n-by-NumDimensions matrix.
Each row represents one embedded point. n is the
number of rows of data X that do not contain
any NaN entries. See Plot Results with NaN Input Data.
Kullback-Leibler divergence between modeled input and output distributions, returned as a nonnegative scalar. For details, see t-SNE Algorithm.
More About
The syntax of a custom distance function is as follows.
function D2 = distfun(ZI,ZJ)tsne passes ZI and ZJ to
your function, and your function computes the distance.
ZIis a 1-by-n vector containing a single row fromX.ZJis an m-by-n matrix containing multiple rows ofX.
Your function returns D2, which is an m-by-1 vector
of distances. The jth element of D2 is the distance
between the observations ZI and ZJ(j,:).
Tip
If your data are not sparse, then usually the built-in distance functions are faster than a function handle.
Algorithms
tsne constructs a set of embedded points in a low-dimensional space whose
relative similarities mimic those of the original high-dimensional points. The embedded points
show the clustering in the original data.
Roughly, the algorithm models the original points as coming from a Gaussian distribution, and the embedded points as coming from a Student’s t distribution. The algorithm tries to minimize the Kullback-Leibler divergence between these two distributions by moving the embedded points.
For details, see t-SNE.
The Distance argument values that begin with fast
(such as "fasteuclidean" and "fastseuclidean")
calculate Euclidean distances using an algorithm that uses extra memory to save
computational time. This algorithm is named "Euclidean Distance Matrix Trick" in Albanie
[1] and elsewhere. Internal
testing shows that this algorithm saves time when the number of predictors is at least 10.
Algorithms starting with fast do not support sparse data.
To find the matrix D of distances between all the points xi and xj, where each xi has n variables, the algorithm computes distance using the final line in the following equations:
The matrix in the last line of the equations is called the Gram matrix. Computing the set of squared distances is faster, but slightly less numerically stable, when you compute and use the Gram matrix instead of computing the squared distances by squaring and summing. For more details, see Albanie [1].
To store the Gram matrix, the software uses a cache with the default size of
1e3 megabytes. You can set the cache size using the
CacheSize name-value argument. If the value of
CacheSize is too large or "maximal", then the
software might try to allocate a Gram matrix that exceeds the available memory. In this
case, the software issues an error.
References
[1] Albanie, Samuel. Euclidean Distance Matrix Trick. June, 2019. Available at https://samuelalbanie.com/files/Euclidean_distance_trick.pdf.
Version History
Introduced in R2017aThe 'fasteuclidean' and
'fastseuclidean' distance metrics accelerate the computation of
Euclidean distances by using a cache and a different algorithm (see Algorithms). Set
the size of the cache using the CacheSize name-value argument.
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)