You are now following this Submission
- You will see updates in your followed content feed
- You may receive emails, depending on your communication preferences
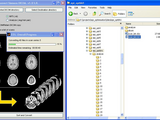
The program offers GUI (dicom_sort_convert_main.m) and .m (example_dscm_call.m) input to select input and output directories and determine options. The main functions are
i) sorting of Siemens MR scanner (IMA) data into directories, removal of patient name from file, and lightly anonymisaton (only PatientName fields but not DOB, gender, PatientID, etc)
ii) Conversion to one-file (.nii) or two-file (.img, .hdr) NIfTI format (one per scan), reformatting more complex scans to use up to six dimensions of the NIfTI structure (time-points, echo, RF channel, phase/magnitude, diffusion gradient, etc). Also writes both DICOM header info and the text section of Siemens header info to a text_header.txt file for each scan.
iii) Generation of a list of all scans performed (scan_list.txt) including most relevant parameters (can easily be customised), making it easy to generate a summary sheet for each subject scanned.
Thanks to Jimmy Shen for sharing, developing and supporting his excellent NIfTI toolbox.
Cite As
Simon Robinson (2026). Siemens DICOM sort and convert to NIfTI (https://www.mathworks.com/matlabcentral/fileexchange/22508-siemens-dicom-sort-and-convert-to-nifti), MATLAB Central File Exchange. Retrieved .
Acknowledgements
Inspired by: Tools for NIfTI and ANALYZE image
General Information
- Version 1.27 (164 KB)
MATLAB Release Compatibility
- Compatible with any release
Platform Compatibility
- Windows
- macOS
- Linux
| Version | Published | Release Notes | Action |
|---|---|---|---|
| 1.27 | see comments in dicom_sort_convert_main.m |
||
| 1.8.0.0 | Please see version changes via 'help dicom_sort_convert_main.m'. |
||
| 1.7.0.0 | please see help |
||
| 1.6.0.0 | Summary field too short to be useful. Use >> help dicom_sort_convert_main |
||
| 1.5.0.0 | The 250 character limit for summaries is too short.
|
||
| 1.4.0.0 | There were a couple of trivial bugs in v1.15 which rendered it a defunct version. I have corrected these. Apologies |
||
| 1.3.0.0 | Splash image (required) was missing from archive |
||
| 1.2.0.0 | v1.14 1) reformats DTI data, 4th dim is grad
|
||
| 1.1.0.0 | v1.14 1) reformats DTI data (4th Dim is grad)
|
||
| 1.0.0.0 |