emukamel/CellSort
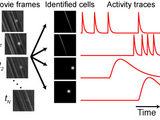
This toolbox includes routines for using principal component analysis (PCA) and independent component analysis (ICA) to extract cellular signals from imaging data sets. A full description and validation of the method is provided in the paper, "Automated Analysis of Cellular Signals from Large-Scale Calcium Imaging Data," Neuron, 63:747 (2009): http://tinyurl.com/cellsort
Cite As
Eran Mukamel (2024). emukamel/CellSort (https://github.com/mukamel-lab/CellSort), GitHub. Retrieved .
MATLAB Release Compatibility
Platform Compatibility
Windows macOS LinuxCategories
Tags
Community Treasure Hunt
Find the treasures in MATLAB Central and discover how the community can help you!
Start Hunting!Discover Live Editor
Create scripts with code, output, and formatted text in a single executable document.
Versions that use the GitHub default branch cannot be downloaded
| Version | Published | Release Notes | |
|---|---|---|---|
| 1.4.0.0 | June 2016: Fixed an inconsistency in CellsortPCA between create_tcov and
|
|
|
| 1.2.0.0 | Fixed a bug affecting performance in recent versions of MATLAB. |
||
| 1.1.0.0 | Fixed a minor typo in documentation for CellsortICA - EAM 11/24/09 |
||
| 1.0.0.0 |