You are now following this Submission
- You will see updates in your followed content feed
- You may receive emails, depending on your communication preferences
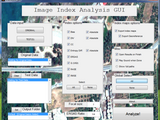
GUI that enables the user to calculate 8 image indices:
1) Bias,
2) CC (Correlation Coefficient),
3) DIV (Difference in Variance)
4) ED (Entropy Difference),
5) ERGAS,
6) UIQI,
7) RASE and
8) RMSE.
The scope of this GUI is to measure the image quality/error between 2
(original/reference image, processed/distorted image) or more images.
The image quality is estimated and quantified by the respective 8 image
indices which are calculated locally (in a neighborhood). User can
specify the size of the neighborhood, by setting the FOCAL SIZE parameter
accordingly.
User can opt to calculate any number of the 8 indices for 2 or more
images (batch mode). So it can be characterized as a batch image index
analysis tool. The index analysis results are written in an Excel file,
named "Focal image analysis.xls" in the same folder. Every index result
is written in its homonymous spreadsheet. If there is no Microsoft Excel
installed in the current computer, the index analysis results are written
in a *.CSV file.
If the user wishes to study and visualize how each index has performed
locally, he can check the button "Export index maps", which results in
outputting every index map for every TEST (processed) image.
Despite the fact that this program has been initially designed in order
to evaluate satellite imagery, all of the 8 above indices can be used for
digital images of any origin (although some of them are specialized for
quality assessment in satellite datasets). Basically, this program is an
enhanced version of this submission:
http://www.mathworks.com/matlabcentral/fileexchange/32637
There are 2 major improvements compared to the above submission:
1) The ability to calculate and evaluate every index locally in the image
2) The form of this program: GUI vs. script (previous)
Usage:
0) Add to path the folder that includes imanalysis_gui function. Also add
all subfolders which are in the same folder with imanalysis_gui.m + fig.
This can be done quickly (and temporarily) by running install_temp.m.
1) Run the GUI by typing imanalysis_gui in Matlab's command window.
2) Load your original (reference) image by clicking the ORIGINAL button.
3) Load your test(s) (processed) image(s) by clicking the TEST(s) button.
4) You can set the output folder, by clicking the 'Set' pushbutton.
However, you may find it useful that by default the program outputs to
the folder containing the original image.
5) Choose which indices you want the program to calculate. You can select
all indices at once, by pushing the 'All' button, under 'Select:' text.
6) You may want to change the FOCAL_SIZE parameter, which as mentioned,
specifies the size of the neighborhood or "window" (filter) in which each
index will be calculated. The larger the focal size, the more the
computations, the more the required time. Usually, we use a neighborhood
of 7 or 9. It is best to input an odd value in this parameter.
7) Optionally, you can check the export index maps button, if you want
the program to output the "index maps" for each index and test image. If
you work with georeferenced imagery, you will probably want to check the
export georeference button as well.
8) Press the 'Analyze!' button.
9) The program performs index analysis with your options. Messages should
appear in the command window, which state its current task. When all
calculations have finished, an Excel (or CSV - read above) file is
created with the results. If you wanted to export the index maps, you can
find them in the newly created folder 'Index maps' (which will be in the
same folder with the GUI).
Important Facts:
1) You can use 2D (grayscale) or 3D (color with 3 or more bands) imagery.
2) Every TEST image must have the exact same resolution and dimensions
with the ORIGINAL image.
3) The ED index is always calculated after conversion to 8bits.
Compatibility:
Oldest version known to have full compatibility: R2011a.
Older versions than R2011a should work if the export index maps option is
off. If you have an older version of Matlab, but want to export the index
maps, then you can download the geotiffwrite function from FEX, and
modify this code accordingly, so it works for you.
Requirements:
Image processing toolbox
Mapping toolbox (geotiffwrite)
Citation:
A. D. Vaiopoulos, "Developing Matlab scripts for image analysis and
quality assessment", Proc. SPIE 8181, 81810B (2011);
Link of the above paper:
http://dx.doi.org/10.1117/12.897806
Cite As
Aristidis Vaiopoulos (2026). Image Quality - Index Analysis GUI (https://www.mathworks.com/matlabcentral/fileexchange/41464-image-quality-index-analysis-gui), MATLAB Central File Exchange. Retrieved .
Acknowledgements
Inspired by: progressbar, dlmcell
General Information
- Version 1.6.0.0 (1020 KB)
MATLAB Release Compatibility
- Compatible with any release
Platform Compatibility
- Windows
- macOS
- Linux
| Version | Published | Release Notes | Action |
|---|---|---|---|
| 1.6.0.0 | -Fixed a bug which was causing incorrect ERGAS values, when user did not opt for index map output (near line 1123). Otherwise, ERGAS values were correct, but the default focal size value was used (=8) (lines 1242,1373). |
||
| 1.4.0.0 | -Fixed a bug (near line 1130). It occured when user opted for rmse, without index maps output. |
||
| 1.3.0.0 | -Improved the code of the GUI function (imanalysis_gui) in order to make it less vulnerable to errors and crashes. |
||
| 1.0.0.0 |