You are now following this Submission
- You will see updates in your followed content feed
- You may receive emails, depending on your communication preferences
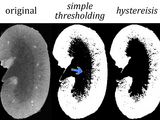
This hysteresis function performs a dual thresholding operation on a grayscale image (2D or 3D) using two threshold values (lower and upper). A trinarisation image is also produced where the lower threshold value is set to 1 and the upper threshold value is set to 2. Hysteresis performs better than standard thresholding (single value) because hysteresis uses a loop to produce a more connect segmentation with fewer isolated pixels. The function help is included below:
Hysteresis3d is a simple function that performs trinarisation and hysteresis for 2D and 3D images. Hysteresis3d was inspired by Peter Kovesi's 2D hysteresis function (http://www.csse.uwa.edu.au/~pk/research/matlabfns/). This 3D function takes advantage of the 3D connectivities of imfill instead of the 2D connectivities of bwselect.
Usage:
[tri,hys]=HYSTERESIS3D(img,t1,t2,conn)
Arguments:
img - image for hysteresis (assumed to be non-negative)
t1 - lower threshold value (fraction b/w 0-1, e.g.: 0.1)
t2 - upper threshold value (fraction b/w 0-1, e.g.: 0.9)
(t1/t2 can be entered in any order, larger one will be set as the upper threshold)
conn - number of connectivities (4 or 8 for 2D; 6, 18, or 26 for 3D)
Returns:
tri - the trinarisation image (values are 0, 1, or 2)
hys - the hysteresis image (logical mask image)
Examples: [tri,hys]=HYSTERESIS3D(img,0.25,0.8,26)
To see an example of hysteresis used to segment a kidney region, please refer to the supplement in QSM of kidney inflammation and fibrosis, NMR Biomed, 2013 Dec;26(12):1853-63 (http://onlinelibrary.wiley.com/doi/10.1002/nbm.3039/abstract). Supplemental material is also available on our CIVMspace: http://www.civm.duhs.duke.edu/lx201204/
Cite As
Luke Xie (2026). Hysteresis thresholding for 3D images (or 2D) (https://www.mathworks.com/matlabcentral/fileexchange/44648-hysteresis-thresholding-for-3d-images-or-2d), MATLAB Central File Exchange. Retrieved .
General Information
- Version 1.0.0.0 (2.19 KB)
MATLAB Release Compatibility
- Compatible with any release
Platform Compatibility
- Windows
- macOS
- Linux
| Version | Published | Release Notes | Action |
|---|---|---|---|
| 1.0.0.0 |