You are now following this Submission
- You will see updates in your followed content feed
- You may receive emails, depending on your communication preferences
This Script is Copyrighted by Tyler L. Coye (2015). Doctoral Candidate in Medicine, Temple University . This method has been downloaded over 6,000 times since its inception.
For those asking if I have formerly published this algorithm, the answer is that I have not due to the time constraints of medical school. However, the significant number of papers using this method is a testament to the value of it in research. I am open to co-authoring this algorithm with someone if they are willing to commit the time to it.
This algorithm is the result of many hours of work and problem solving. If you use this algorithm in your work, please copy the following citation:
* * * * *
Coye, Tyler (2015). A Novel Retinal Blood Vessel Segmentation Algorithm for Fundus Images (http://www.mathworks.com/matlabcentral/fileexchange/50839), MATLAB Central File Exchange.[Month and Day Retrieved], [Year Retrieved].
* * * * *
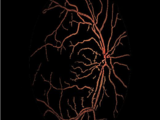
This Script segments retinal blood vessels in a fundus image, which is a difficult challenge to overcome.
Included in the .zip file are the main file, labeled 'CoyeFilter.m' as well as the overlay algorithm and Isodata algorithm. The 'CoyeFilter.m' utilizes both overlay and isodata methods.
In the .zip is a sample set of fundus images for you to test the Script on.
Instead of using the green channel, as many papers have commonly done in the past, I converted the RGB to Gray using PCA through a process I developed in my algorithm " Hybrid Lesion Detection Algorithm Using Principle Component Analysis."
* * * * * *
For references to other papers using this method see: https://scholar.google.com/scholar?oi=bibs&hl=en&cites=9803621023695796972&as_sdt=5
Examples of some of the papers which have utilized this method for segmenting blood vessels in the retina:
Bandara, A. M. R. R., and P. W. G. R. M. P. B. Giragama. "A retinal image enhancement technique for blood vessel segmentation algorithm." Industrial and Information Systems (ICIIS), 2017 IEEE International Conference on. IEEE, 2017.
Buglione, John. Use of Compton Backscatter X-ray Imaging in Agriculture. ME thesis. University of Pennsylvania, Philadelphia, 2016. http://www.jakebuglione.com/thesis.pdf
Furtado, Pedro, et al. "Segmentation of Eye Fundus Images by density clustering in diabetic retinopathy." Biomedical & Health Informatics (BHI), 2017 IEEE EMBS International Conference on. IEEE, 2017.
Gu, Lin, et al. "Semi-supervised learning for biomedical image segmentation via forest oriented super pixels (voxels)." International Conference on Medical Image Computing and Computer-Assisted Intervention. Springer, Cham, 2017.
Intaramanee, Tanin, et al. "Optic disc detection via blood vessels origin using Morphological end point." Advanced Informatics: Concepts, Theory And Application (ICAICTA), 2016 International Conference On. IEEE, 2016.
Saponaro, P., et al. "Three-Dimensional Segmentation of Vesicular Networks of Fungal Hyphae in Macroscopic Microscopy Image Stacks." arXiv preprint arXiv:1704.02356 (2017).
Saranya, M., and A. Grace Selvarani. "Fundus Image Screening for Diabetic Retinopathy." Indian Journal of Science and Technology 9.25 (2016).
Travassos, C., et al. "DENSITY SEGMENTATION IN DETECTION OF DIABETIC RETINOPATHY." (2017).
* * * * * *
The Script for the Segmentation algorithm is below:
%% Computer Assisted Retinal Blood Vessel Segmentation Algorithm
% Developed and Copyrighted by Tyler L. Coye (2015)
%
% Read Image
I = imread('13_right.jpeg')
% Resize image for easier computation
B = imresize(I, [584 565])
% Read image
im = im2double(B);
% Convert RGB to Gray via PCA
lab = rgb2lab(im);
f = 0;
wlab = reshape(bsxfun(@times,cat(3,1-f,f/2,f/2),lab),[],3);
[C,S] = pca(wlab);
S = reshape(S,size(lab));
S = S(:,:,1);
gray = (S-min(S(:)))./(max(S(:))-min(S(:)));
%% Contrast Enhancment of gray image using CLAHE
J = adapthisteq(gray,'numTiles',[8 8],'nBins',128);
%% Background Exclusion
% Apply Average Filter
h = fspecial('average', [9 9]);
JF = imfilter(J, h);
figure, imshow(JF)
% Take the difference between the gray image and Average Filter
Z = imsubtract(JF, J);
figure, imshow(Z)
%% Threshold using the IsoData Method
level=isodata(Z) % this is our threshold level
%% Convert to Binary
BW = im2bw(Z, level-.008)
%% Remove small pixels
BW2 = bwareaopen(BW, 100)
%% Overlay
BW2 = imcomplement(BW2)
out = imoverlay(B, BW2, [0 0 0])
figure, imshow(out)
Cite As
Tyler Coye (2026). Novel Retinal Vessel Segmentation Algorithm: Fundus Images (https://www.mathworks.com/matlabcentral/fileexchange/50839-novel-retinal-vessel-segmentation-algorithm-fundus-images), MATLAB Central File Exchange. Retrieved .
General Information
- Version 2.0.0.2 (10.2 MB)
MATLAB Release Compatibility
- Compatible with any release
Platform Compatibility
- Windows
- macOS
- Linux
| Version | Published | Release Notes | Action |
|---|---|---|---|
| 2.0.0.2 | Updated Title previously: A novel retinal blood vessel segmentation algorithm for fundus images. |
||
| 2.0.0.1 | Updated description and edited citation |
||
| 2.0.0.0 | updated citations.
|
||
| 1.0.0.0 | Minor description changes
|