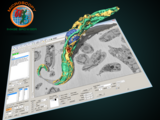
MIB is a package for segmentation of multi-dimensional (2D-4D) microscopy datasets
You are now following this Submission
- You will see updates in your followed content feed
- You may receive emails, depending on your communication preferences
PLEASE NOTE!
This is an older version tree of MIB, the newer version for Matlab R2014b and newer available from here
http://se.mathworks.com/matlabcentral/fileexchange/63402-microscopy-image-browser-2--mib2-
With MIB you can analyse, segment and visualize various multidimensional datasets from both light and electron microscopy.
See more further details and tutorials on MIB website: http://mib.helsinki.fi/index.html
I would like to acknowledge Matlab File Exchange user community and especially the authors whose functions were utilized during development of the program:
http://mib.helsinki.fi/acknowledgements.html
Cite As
Ilya Belevich (2026). Microscopy Image Browser (MIB) (https://github.com/Ajaxels/MIB), GitHub. Retrieved .
Acknowledgements
Inspired by: view3d.m, Local Normalization, extrema.m, extrema2.m, IMAGEVIEWER, findjobj - find java handles of Matlab graphic objects, 3D Euclidean Distance Transform for Variable Data Aspect Ratio, Region Adjacency Graph (RAG), stlwrite - write ASCII or Binary STL files, maxflow, Viewer3D, export_fig, Hessian based Frangi Vesselness filter, Accurate Fast Marching, Fast/Robust Template Matching, imgaussian, Image Edge Enhancing Coherence Filter Toolbox, Highly portable JSON-input parser, Jann5s/measuretool, Render RGB text over RGB or Grayscale Image, improved xlswrite.m, xml2struct, struct2xml, IMCLIPBOARD, Hardware accelerated 3D viewer for MATLAB, mdrohmann/mtocpp, drifty_shifty_deluxe.m, regionprops3
General Information
- Version 1.302.0.0 (41.1 MB)
-
View License on GitHub
MATLAB Release Compatibility
- Compatible with any release
Platform Compatibility
- Windows
- macOS
- Linux
Versions that use the GitHub default branch cannot be downloaded
| Version | Published | Release Notes | Action |
|---|---|---|---|
| 1.302.0.0 | 1.302 Improved and simplified the segmentation panel
|
||
| 1.23.0.0 | new image alignment; IMOD enhanced compatibility, improvements and bug fixes
|
||
| 1.2.0.0 | Version 1.20 20.06.2016 (volume rendering/new installer) |
||
| 1.1.0.0 | updated description |