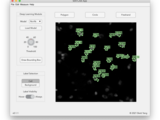
A deep learning-enabled cell fluorescence measurement tool
https://archive.brettyang.au/neuroscience/computation/2021/08/21/CTCF-ML/
You are now following this Submission
- You will see updates in your followed content feed
- You may receive emails, depending on your communication preferences
This ROI labeller was designed for the purpose of cell fluorescence measurements, though its functionalities are not limited to this purpose. One may export region of interests data (MATLAB cell array of ROI objects), as a binary mask, or as an instance mask.
A deep learning-based segmentation model is used to speed up the labelling process for cellular objects.
Created for the Laboratory of Molecular Neuroscience and Dementia, University of Sydney.
Cite As
Zeyi Yang (2026). Semi-automated Cell Fluorescence Measurements (https://github.com/where-is-brett/cell-fluorescence-ml/releases/tag/v0.1.1), GitHub. Retrieved .
General Information
- Version 0.1.1 (127 KB)
-
View License on GitHub
MATLAB Release Compatibility
- Compatible with any release
Platform Compatibility
- Windows
- macOS
- Linux
| Version | Published | Release Notes | Action |
|---|---|---|---|
| 0.1.1 | See release notes for this release on GitHub: https://github.com/where-is-brett/cell-fluorescence-ml/releases/tag/v0.1.1 |
To view or report issues in this GitHub add-on, visit the GitHub Repository.
To view or report issues in this GitHub add-on, visit the GitHub Repository.