anova
Description
An anova object contains the results of a one-, two-, or N-way
ANOVA. Use the properties of an anova object to determine if the
means in a set of response data differ with respect to the values (levels) of a factor or
multiple factors. The object properties include information about the coefficient estimates,
ANOVA model fit to the response data, and factors used to perform the analysis.
Creation
Syntax
Description
aov = anova(y)anova object aov for
the response data in the matrix y. Each column of
y is treated as a different factor value.
aov = anova(tbl,responseVarName)tbl as factors and response data. The
responseVarName argument specifies which variable contains the
response data.
aov = anova(tbl,formula)formula use only the variable names in
tbl.
aov = anova(___,Name=Value)
Input Arguments
y — Response data
matrix | numeric vector
Response data, specified as a matrix or a numeric vector.
If
yis a matrix,anovatreats each column ofyas a separate factor value in a one-way ANOVA. In this design, the function evaluates whether the population means of the columns are equal. Use this design when you want to perform a one-way ANOVA on data that is equally divided between each group (balanced ANOVA).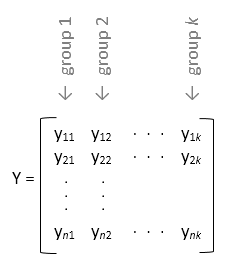
If
yis a numeric vector, you must also specify either thefactorsortblinput argument. For a one-way ANOVA,factorsis a cell array of character vectors or a vector in which each element represents the factor value of the corresponding element iny.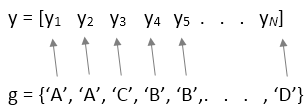
For an N-way ANOVA,
factorsis a cell array of vectors in which each cell is treated as a separate factor. Alternatively, for an N-way ANOVA, you can provide a tabletblin which each variable is treated as a separate factor. Use this design when you want to perform a two- or N-way ANOVA, or when factor values correspond to different numbers of observations iny(unbalanced ANOVA).
Note
The anova function ignores NaN
values, <undefined> values, empty characters, and empty
strings in y. If factors or
tbl contains NaN or
<undefined> values, or empty characters or strings, the
function ignores the corresponding observations in y. The ANOVA
is balanced if each factor value has the same number of observations after the
function disregards empty or NaN values. Otherwise, the function
performs an unbalanced ANOVA.
Data Types: single | double
factors — factors and factor values
numeric vector | logical vector | categorical vector | string vector | character vector | cell array of vectors
Factors and factor values for the ANOVA, specified as a numeric, logical, categorical, string, or character vector, or a cell array of vectors. Factors and factor values are sometimes called grouping variables and group names, respectively.
For a one-way ANOVA, factors is a vector or cell array of
character vectors in which each element represents the factor value of the observation
in y at the same position. The anova
function groups observations in y by their factor values during
the ANOVA. The length of factors must be the same as the length
of y.
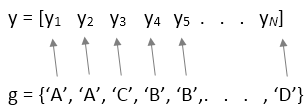
For a two- or N-way ANOVA, factors is a cell array of vectors
in which each cell corresponds to a different factor. Each vector contains the values
of the corresponding factor and must have the same length as y.
Factor values are associated with observations in y by their
index.
If factors contains NaN values,
anova ignores the corresponding observations in
y.
For more information on factors, see Grouping Variables.
Note
If factors or tbl contains
NaN values, <undefined> values, empty
characters, or empty strings, the anova function ignores the
corresponding observations in y. The ANOVA is balanced if each factor
value has the same number of observations after the function disregards empty or
NaN values. Otherwise, the function performs an unbalanced
ANOVA.
Example: [1,2,1,3,1,...,3,1]
Example: ["white","red","white",...,"black","red"]
Example: school=["Springfield","Springfield","Springfield","Arlington","Springfield","Arlington","Arlington"];
monthnumber=[6,12,1,9,4,6,2];
factors={school,monthnumber};
Data Types: single | double | logical | categorical | char | string | cell
tbl — Factors, factor values, and response data
table
Factors, factor values, and response data, specified as a table. The variables of
tbl can contain numeric, logical, categorical, character
vector, or string elements, or cell arrays of characters. When you specify
tbl, you must also specify the response data
y, responseVarName, or
formula.
If you specify the response data in
y, the table variables represent only the factors for the ANOVA. A factor value in a variable oftblcorresponds to the observation inyat the same position.tblmust have the same number of rows as the length ofy. IftblcontainsNaNvalues, thenanovaignores the corresponding observations iny.If you do not specify
y, you must indicate which variable intblcontains the response data by using theresponseVarNameorformulainput argument. You can also choose a subset of factors intblto use in the ANOVA by setting the name-value argumentFactorNames. Theanovafunction associates the values of the factor variables intblwith the response data in the same row.
Note
If factors or tbl contains
NaN values, <undefined> values, empty
characters, or empty strings, the anova function ignores the
corresponding observations in y. The ANOVA is balanced if each factor
value has the same number of observations after the function disregards empty or
NaN values. Otherwise, the function performs an unbalanced
ANOVA.
Example: mountain=table(altitude,temperature,soilpH);
anova(mountain,"soilpH")
Data Types: table
responseVarName — Name of response data
string scalar | character vector
Name of the response data, specified as a string scalar or character vector.
responseVarName indicates which variable in
tbl contains the response data. When you specify
responseVarName, you must also specify the
tbl input argument.
Example: "r"
Data Types: char | string
formula — ANOVA model
string scalar | character vector
ANOVA model, specified as a string scalar or a character vector in Wilkinson notation. anova supports the use of
parentheses and commas to specify nested factors in formula. For
example, you can specify that factor f1 is nested inside factor
f2 by including the term f1(f2) in
formula. To specify that f1 is nested inside
two factors, f2 and f3, include the term
f1(f2,f3). When you specify formula, you
must also specify tbl.
Example: "r ~ f1 + f2 + f3 + f1:f2:f3"
Example: "MPG ~ Origin + Model(Origin)"
Data Types: char | string
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: anova(factors,y,CategoricalFactors=[1 2],FactorNames=["school"
"major" "age"],ResponseName="GPA") specifies the first two factors in
factors as categorical, the factor names as
"school", "major", and "age",
and the name of the response variable as "GPA".
CategoricalFactors — Factors to treat as categorical
"all" (default) | numeric vector | logical vector | string vector | cell array of character vectors
Factors to treat as categorical, specified as a numeric, logical, or string
vector, or a cell array of character vectors. When
CategoricalFactors is set to the default value
"all", the anova function treats all
factors as categorical.
Specify CategoricalFactors as one of the following:
A numeric vector with indices between 1 and N, where N is the number of factor variables. The
anovafunction treats factors with indices inCategoricalFactorsas categorical. The index of a factor is the order in which it appears in the columns of matrixy, the cells offactors, or the columns oftbl.A logical vector of length N, where a
trueentry means that the corresponding factor is categorical.A string vector or cell array of factor names. The factor names must match the names in
tblorFactorNames.
Example: CategoricalFactors=["Location"
"Smoker"]
Example: CategoricalFactors=[1 3 4]
Data Types: single | double | logical | char | string | cell
FactorNames — Factor names
string vector | cell array of character vectors
Factor names, specified as a string vector or a cell array of character vectors.
If you specify
tblin the call toanova,FactorNamesmust be a subset of the table variables intbl.anovauses only the factors specified inFactorNames. In this case, the default value ofFactorNamesis the collection of names of the factor variables intbl.If you specify the matrix
yorfactorsin the call toanova, you can specify any names forFactorNames. In this case, the default value ofFactorNamesis["Factor1","Factor2",…,"FactorN"], where N is the number of factors.
When you specify formula, anova
ignores FactorNames.
Example: FactorNames=["time","latitude"]
Data Types: char | string | cell
ModelSpecification — Type of ANOVA model to fit
"linear" (default) | "interactions" | "purequadratic" | "quadratic" | "polyIJK" | "full" | integer | string scalar | character vector | terms matrix
Type of ANOVA model to fit, specified as one of the options in the following
table or an integer, string scalar, character vector, or terms matrix. The default
value for ModelSpecification is
"linear".
| Option | Terms Included in ANOVA Model |
|---|---|
"linear" (default) | Main effect (linear) terms |
"interactions" | Main effect and pairwise interaction terms |
"purequadratic" | Main effects and squared main effects. All factors must be continuous
to use this option. Set CategoricalFactors = [] to
specify all factors as continuous. |
"quadratic" | Main effects, squared main effects, and pairwise interaction terms. All factors must be continuous to use this option. |
"polyIJK" | Polynomial terms up to degree I for the first factor, degree J for the second factor, and so on. The degree of an interaction term cannot exceed the maximum exponent of a main term. You must specify a degree for each factor. |
"full" | Main effect and all interaction terms |
To include all main effects and interaction levels up to the
kth level, set ModelSpecification equal to
k. When ModelSpecification is an integer,
the maximum level of an interaction term in the ANOVA model is the minimum between
ModelSpecification and the number of factors.
If you specify formula, anova
ignores ModelSpecification.
You can also specify the terms of an ANOVA regression model using one of the following:
Double or single terms matrix, T, with a column for each factor. Each term in the ANOVA model is a product corresponding to a row of T. The row elements are the exponents of their corresponding factors. For example,
T(i,:) = [1 2 1]means that termiis . Because theanovafunction automatically includes a constant term in the ANOVA model, you do not need to include a row of zeros in the terms matrix.Character vector or string scalar formula in Wilkinson notation, representing one or more terms.
anovasupports the use of parentheses and commas to specify nested factors, as described informula. The formula must use names contained inFactorNames,ResponseName, or table variable names iftblis specified.
Example: ModelSpecification="poly3212"
Example: ModelSpecification=3
Example: ModelSpecification="r ~ c1*c2"
Example: ModelSpecification=[0 0 0;1 0 0;0 1 0;0 0
1]
Data Types: single | double | char | string
RandomFactors — Factors to treat as random
"all" | numeric vector | logical vector | string vector | cell array of character vectors
Factors to treat as random rather than fixed, specified as a numeric, logical,
or string vector, or a cell array of character vectors. The
anova function treats an interaction term as random if
it contains at least one random factor. The default value is [],
meaning all factors are fixed. To specify all factors as random, set
RandomFactors to "all".
Specify RandomFactors as one of the following:
A numeric vector with indices between 1 and N, where N is the number of factor variables. The
anovafunction treats factors with indices inRandomFactorsas random. The index of a factor is the order in which it appears in the columns of matrixy, the cells offactors, or the columns oftbl.A logical vector of length N, where a
trueentry means that the corresponding factor is random.A string vector or cell array of factor names. The factor names must match the names in
tblorFactorNames.
Example: RandomFactors=[1]
Example: RandomFactors=[1 0 0]
Data Types: single | double | logical | char | string | cell
ResponseName — Name of response variable
string scalar | character vector
Name of the response variable, specified as a string scalar or a character
vector. If you specify responseVarName or
formula, anova ignores
ResponseName.
Example: ResponseName="soilpH"
Data Types: char | string
SumOfSquaresType — Type of sum of squares
"three" (default) | "two" | "one" | "hierarchical"
Type of sum of squares used to perform the ANOVA, specified as "three",
"two", "one", or
"hierarchical". For a model containing main effects but no
interactions, the value of SumOfSquaresType influences the
computations on the unbalanced data only.
The sum of squares of a term () is defined as the reduction in the sum of squares error (SSE) obtained by adding the term to a model that excludes it. The formula for the sum of squares of a term Term has the form
where n is the number of observations, are the response data, are the factors used to perform the ANOVA, is a model that excludes Term, and is a model that includes Term. Both and are specified by SumOfSquaresType. The variables and are the sum of squares errors for and , respectively. You can specify and using one of the options for SumOfSquaresType described
in the following table.
| Option | Type of Sum of Squares |
|---|---|
"three" (default) | is the full ANOVA model specified in the property
|
"two" | is a model composed of all terms in the ANOVA model
specified in the property |
"one" | is a model composed of all the terms that precede
Term in the ANOVA model specified in the property
|
"hierarchical" | and are defined as in Type II, except powers of Term are treated as terms that contain Term. |
Example: SumOfSquaresType="hierarchical"
Data Types: char | string
Properties
CategoricalFactors — Indices of categorical factors
numeric vector
This property is read-only.
Indices of categorical factors, specified as a numeric vector. This property is set
by the CategoricalFactors name-value argument.
Data Types: double
Coefficients — Fitted ANOVA model coefficients
double vector
This property is read-only.
Fitted ANOVA model coefficients, specified as a double vector. The
anova function expands each categorical factor into
F dummy variables, where F is the number of
values for the factor. Each dummy variable is fit with a different coefficient during
the ANOVA. Continuous factors have coefficients that are constant across factor values.
For example, let y be a set of response data and
factor1 be a continuous factor. Let factor2 be a
categorical factor with values value1, value2, and
value3. The formula "y ~ 1 + factor1 + factor2"
expands to "y ~ 1 + factor1 + (factor2==value1) + (factor2==value2) +
(factor2==value3)" and anova fits the expanded
formula with coefficients.
Data Types: single | double
ExpandedFactorNames — Names of coefficients
string vector
This property is read-only
Names of coefficients, specified as a string vector of names. The
anova function expands each categorical factor into
F dummy variables, where F is the number of
values for the factor. The vector ExpandedFactorNames contains the
name of each dummy variable. For more information, see Coefficients.
Data Types: string
FactorNames — Names of factors
string vector
This property is read-only.
Names of the factors used to fit the ANOVA model, specified as a string vector of
names. This property is set by the tbl input argument or the
FactorNames name-value argument.
Data Types: string
Factors — Names and values of factors
table
This property is read-only.
Names and values of the factors used to fit the ANOVA model, specified as a table.
The names of the table variables are the factor names, and each variable contains the
values of its corresponding factor. If the factors used to fit the model are not given
as a table, anova converts them into a table with one column per
factor.
This property is set by one of the following:
tblinput argumentMatrix
yinput argument together with theFactorNamesname-value argumentVector
yinput argument together with thefactorsinput argument and theFactorNamesname-value argument
Data Types: table
Formula — ANOVA model
LinearFormulaWithNesting object
This property is read-only.
ANOVA model, specified as a LinearFormulaWithNesting object. This
property is set by the formula input argument or the
ModelSpecification name-value argument.
Metrics — Model metrics
table
Model metrics, specified as a table. The table Metrics has
these variables:
MSE — Mean squared error.
RMSE — Root mean squared error, which is the square root of MSE.
SSE — Sum of squares of the error.
SSR — Sum of squares regression.
SST — Total sum of squares.
RSquared — Coefficient of determination, also known as .
AdjustedRSquared — value, adjusted for the number of coefficients. This value is given by the formula , where n is the number of observations, and p is the number of coefficients. A higher value for indicates a better fit for the ANOVA model.
Data Types: table
NumObservations — Number of observations
positive integer
This property is read-only.
Number of observations used to fit the ANOVA model, specified as a positive integer.
Data Types: double
RandomFactors — Indices of random factors
numeric vector
This property is read-only.
Indices of random factors, specified as a numeric vector. This property is set by
the RandomFactors name-value argument.
Data Types: double
Residuals — Residual values
n-by-2 table
This property is read-only.
Residual values, specified as an n-by-2 table, where
n is the number of observations. Residuals has
two variables:
Raw contains the observed minus fitted values.
Pearson contains the raw residuals divided by the root mean squared error (RMSE).
Data Types: table
SumOfSquaresType — Type of sum of squares
"three" (default) | "two" | "one" | "hierarchical"
This property is read only.
Type of sum of squares used when fitting the ANOVA model, specified as "three",
"two", "one", or "hierarchical". This property is set by the
SumOfSquaresType name-value argument.
Data Types: string
ResponseName — Name of response variable
string scalar | character vector
This property is read-only.
Name of the response variable, specified as a string scalar or character vector.
This property is set by the responseVarName input argument or the
ResponseName name-value argument.
Data Types: char | string
Y — Response data
numeric vector
This property is read-only.
Response data used to fit the ANOVA model, specified as a numeric vector. This
property is set by the y input argument, or the
tbl input argument together with the
responseVarName input argument.
Data Types: single | double
Object Functions
boxchart | Box chart (box plot) for analysis of variance (ANOVA) |
groupmeans | Mean response estimates for analysis of variance (ANOVA) |
multcompare | Multiple comparison of means for analysis of variance (ANOVA) |
plotComparisons | Interactive plot of multiple comparisons of means for analysis of variance (ANOVA) |
stats | Analysis of variance (ANOVA) table |
varianceComponent | Variance component estimates for analysis of variance (ANOVA) |
Examples
Perform One-Way ANOVA for Matrix Data
Load popcorn yield data.
load popcorn.mat The columns of the 6-by-3 matrix popcorn contain popcorn yield observations in cups for three different brands. Perform a one-way ANOVA to test the null hypothesis that the popcorn yield is not affected by the brand of popcorn.
aov = anova(popcorn)
aov =
1-way anova, constrained (Type III) sums of squares.
Y ~ 1 + Factor1
SumOfSquares DF MeanSquares F pValue
____________ __ ___________ ____ __________
Factor1 15.75 2 7.875 18.9 7.9603e-05
Error 6.25 15 0.41667
Total 22 17
Properties, Methods
aov is an anova object that contains the results of the one-way ANOVA.
The Factor1 row of the ANOVA table shows statistics for the model term Factor1, and the Error row shows statistics for the entire model. The sum of squares and the degrees of freedom are given in the SumOfSquares and DF columns, respectively. The Total degrees of freedom is the total number of observations minus one, which is 18 – 1 = 17. The Factor1 degrees of freedom is the number of factor values minus one, which is 3 – 1 = 2. The Error degrees of freedom is the total degrees of freedom minus the Factor1 degrees of freedom, which is 17 – 2 = 15.
The mean squares, given in the MeanSquares column, are calculated with the formula SumOfSquares/DF. The F-statistic is the ratio of the mean squares, which is 7.875/0.41667 = 18.9. The F-statistic follows an F-distribution with degrees of freedom 2 and 15. The p-value is calculated using the cumulative distribution function (cdf). The p-value for the F-statistic is small enough that the null hypothesis can be rejected at the 0.01 significance level. Therefore, the brand of popcorn has a significant effect on the popcorn yield.
Perform Two-Way ANOVA for Vector Data
Load popcorn yield data.
load popcorn.matThe columns of the 6-by-3 matrix popcorn contain popcorn yield observations in cups for the brands Gourmet, National, and Generic. The first three rows of the matrix correspond to popcorn that was popped with an oil popper, and the last three rows correspond to popcorn that was popped with an air popper.
Create string vectors containing factor values for the brand and popper type. Use the function repmat to repeat copies of strings.
brand = [repmat("Gourmet",6,1);repmat("National",6,1);repmat("Generic",6,1)]; poppertype = [repmat("Air",3,1);repmat("Oil",3,1);repmat("Air",3,1);repmat("Oil",3,1);repmat("Air",3,1);repmat("Oil",3,1)]; factors = {brand,poppertype};
Perform a two-way ANOVA to test the null hypothesis that the popcorn yield is not affected by the brand of popcorn or the type of popper.
aov = anova(factors,popcorn(:),FactorNames=["Brand" "PopperType"])
aov =
2-way anova, constrained (Type III) sums of squares.
Y ~ 1 + Brand + PopperType
SumOfSquares DF MeanSquares F pValue
____________ __ ___________ ___ __________
Brand 15.75 2 7.875 63 1e-07
PopperType 4.5 1 4.5 36 3.2548e-05
Error 1.75 14 0.125
Total 22 17
Properties, Methods
aov is an anova object containing the results of the two-way ANOVA. The small p-values indicate that both the brand and popper type have a statistically significant effect on the popcorn yield.
Compute the mean response estimates to see which brand and popper type produce the most popcorn.
groupmeans(aov,["Brand" "PopperType"])
ans=6×6 table
Brand PopperType Mean SE MeanLower MeanUpper
__________ __________ ____ _______ _________ _________
"Gourmet" "Air" 5.75 0.16667 5.0329 6.4671
"National" "Air" 4.25 0.16667 3.5329 4.9671
"Generic" "Air" 3.5 0.16667 2.7829 4.2171
"Gourmet" "Oil" 6.75 0.16667 6.0329 7.4671
"National" "Oil" 5.25 0.16667 4.5329 5.9671
"Generic" "Oil" 4.5 0.16667 3.7829 5.2171
The table shows the mean response estimates with their standard error and 95% confidence bounds. The mean response estimates indicate that the Gourmet brand popped in an oil popper yields the most popcorn.
Perform Two-Way ANOVA with Random Effects
Load the patient sample data.
load patients.matCreate a table of factors from the Age and Smoker variables.
tbl = table(Age,Smoker,VariableNames=["Age" "SmokingStatus"]);
The factor SmokingStatus is a randomly sampled categorical factor, and Age is a continuous factor. Perform a two-way ANOVA to test the null hypothesis that systolic blood pressure is not affected by age or smoking status.
aov = anova(tbl,Systolic,CategoricalFactors=2,RandomFactors=2)
aov =
2-way anova, constrained (Type III) sums of squares.
Y ~ 1 + Age + SmokingStatus
SumOfSquares DF MeanSquares F pValue
____________ __ ___________ ______ __________
Age 37.562 1 37.562 1.6577 0.20098
SmokingStatus 2182.9 1 2182.9 96.337 3.3613e-16
Error 2198 97 22.659
Total 4461.2 99
Properties, Methods
aov is an anova object that contains the results of the two-way ANOVA. The p-value for Age is larger than 0.05. At the 95% confidence level, not enough evidence exists to reject the null hypothesis that age does not have a statistically significant effect on systolic blood pressure. SmokingStatus has a p-value smaller than 0.05, indicating that smoking status has a statistically significant effect on systolic blood pressure.
To investigate whether the variability of the random factor SmokingStatus has an effect on the SmokingStatus mean square, use the object functions varianceComponent and stats.
v = varianceComponent(aov)
v=2×3 table
VarianceComponent VarianceComponentLower VarianceComponentUpper
_________________ ______________________ ______________________
SmokingStatus 48.31 9.0308 49707
Error 22.659 17.425 30.68
[~,ems] = stats(aov)
ems=3×5 table
Type ExpectedMeanSquares MeanSquaresDenominator DFDenominator FDenominator
________ ___________________________________ ______________________ _____________ ____________
Age "fixed" "5135.47*Q(Age)+V(Error)" 22.659 97 MS(Error)
SmokingStatus "random" "44.7172*V(SmokingStatus)+V(Error)" 22.659 97 MS(Error)
Error "random" "V(Error)"
Inserting the VarianceComponent values into the SmokingStatus formula for ExpectedMeanSquares gives 44.7172*48.3098+22.6594 = 2.1829e+03. To see how much the variance component of SmokingStatus affects the expected mean squares, divide the SmokingStatus term of ExpectedMeanSquares by ExpectedMeanSquares to get 44.7172*48.3098/2.1829e+03 = 0.9896. This calculation shows that the SmokingStatus variance component contributes to almost 99% of the SmokingStatus expected mean squares.
Perform ANOVA for Data in Table
Load data of the results for five exams taken by 120 students.
load examgrades.matCreate a table with variables for the math, biology, history, literature, and multi-subject comprehensive exams.
subject = ["math" "biology" "history" "literature" "comprehensive"]; grades = table(grades(:,1),grades(:,2),grades(:,3),grades(:,4),grades(:,5),VariableNames=subject)
grades=120×5 table
math biology history literature comprehensive
____ _______ _______ __________ _____________
65 77 69 75 69
61 74 70 66 68
81 80 71 74 79
88 76 80 88 79
69 77 74 69 76
89 93 78 77 80
55 64 60 50 63
84 83 80 77 78
86 75 81 87 79
84 82 86 92 85
71 70 73 81 79
81 88 80 79 83
84 78 80 74 80
81 77 81 83 79
78 66 90 84 75
67 74 73 76 72
⋮
Perform a four-way ANOVA for the continuous factors math, biology, history, and literature, and the response data comprehensive.
aov = anova(grades,"comprehensive",CategoricalFactors = [])aov =
N-way anova, constrained (Type III) sums of squares.
comprehensive ~ 1 + math + biology + history + literature
SumOfSquares DF MeanSquares F pValue
____________ ___ ___________ ______ __________
math 58.973 1 58.973 6.1964 0.014231
biology 100.35 1 100.35 10.544 0.0015275
history 243.89 1 243.89 25.626 1.5901e-06
literature 152.22 1 152.22 15.994 0.00011269
Error 1094.5 115 9.5173
Total 3291 119
Properties, Methods
aov is an anova object that contains the results of the four-way ANOVA. The p-values of all factors are all smaller than 0.05, indicating that each subject exam can be used to predict a student's grade on the comprehensive exam. Display the estimated coefficients of the ANOVA model.
coef = aov.Coefficients
coef = 5×1
21.9901
0.0997
0.1805
0.2563
0.1701
The coefficient corresponding to the history exam is the largest; therefore, history makes the largest contribution to the predicted value of comprehensive.
Compare Two anova Objects Created Using Table
Load popcorn yield data.
load popcorn.matThe columns of the 6-by-3 matrix popcorn contain popcorn yield observations for the brands Gourmet, National, and Generic. The first three rows of the matrix correspond to popcorn that was popped with an oil popper, and the last three rows correspond to popcorn that was popped with an air popper.
Create a table containing variables representing the brand, popper type, and popcorn yield by using the repmat and table functions.
brand = [repmat("Gourmet",6,1);repmat("National",6,1);repmat("Generic",6,1)]; poppertype = [repmat("air",3,1);repmat("oil",3,1);repmat("air",3,1);repmat("oil",3,1);repmat("air",3,1);repmat("oil",3,1)]; tbl = table(brand,poppertype,popcorn(:),VariableNames=["Brand" "PopperType" "PopcornYield"]);
Perform a two-way ANOVA to test the null hypothesis that the popcorn yield is the same across the three brands and the two popper types. Specify the ANOVA model formula using Wilkinson notation.
aovLinear = anova(tbl,"PopcornYield ~ Brand + PopperType")aovLinear =
2-way anova, constrained (Type III) sums of squares.
PopcornYield ~ 1 + Brand + PopperType
SumOfSquares DF MeanSquares F pValue
____________ __ ___________ ___ __________
Brand 15.75 2 7.875 63 1e-07
PopperType 4.5 1 4.5 36 3.2548e-05
Error 1.75 14 0.125
Total 22 17
Properties, Methods
aovLinear is an anova object that contains the results of the two-way ANOVA. The ANOVA model for aovLinear is linear and does not include an interaction term. The small p-values indicate that both the brand and popper type have a significant effect on the popcorn yield.
To investigate whether the interaction between the brand and popper type has a significant effect on the popcorn yield, perform a two-way ANOVA with a model that contains the interaction term Brand:PopperType.
aovInteraction = anova(tbl,"PopcornYield ~ Brand + PopperType + Brand:PopperType")aovInteraction =
2-way anova, constrained (Type III) sums of squares.
PopcornYield ~ 1 + Brand*PopperType
SumOfSquares DF MeanSquares F pValue
____________ __ ___________ ____ __________
Brand 15.75 2 7.875 56.7 7.679e-07
PopperType 4.5 1 4.5 32.4 0.00010037
Brand:PopperType 0.083333 2 0.041667 0.3 0.74622
Error 1.6667 12 0.13889
Total 22 17
Properties, Methods
The ANOVA model for the anova object aovInteraction includes the interaction term Brand:PopperType. The p-value for the Brand:PopperType term is larger than 0.05. Therefore, not enough evidence exists to conclude that the brand and popper type have an interaction effect on the popcorn yield.
The Metrics property of an anova object provides statistics about the fit of the ANOVA model. To determine which model is a better fit for the response data, display the Metrics property of aovLinear and aovInteraction.
aovLinear.Metrics
ans=1×7 table
MSE RMSE SSE SSR SST RSquared AdjustedRSquared
_____ _______ ____ _____ ___ ________ ________________
0.125 0.35355 1.75 20.25 22 0.92045 0.88731
aovInteraction.Metrics
ans=1×7 table
MSE RMSE SSE SSR SST RSquared AdjustedRSquared
_______ _______ ______ ______ ___ ________ ________________
0.13889 0.37268 1.6667 20.333 22 0.92424 0.78535
The metrics tables show that the mean squared error (MSE) is slightly smaller for the linear model than for the interaction model. The adjusted R-squared value is higher for the linear model. Together, these metrics suggest that the linear model is a better fit for the popcorn data than the interaction model.
Perform Nested Two-Way ANOVA
Load the sample car data.
load carbig.matThe variable Model contains data for the car model, and the variable Origin contains data for the country in which the car is manufactured. Convert Model and Origin from character arrays with trailing whitespace to string vectors.
Model = strtrim(string(Model)); Origin = strtrim(string(Origin));
The variable MPG contains mileage data for the cars. Create a table containing data for the model, country of origin, and mileage of the cars manufactured in Japan and the United States.
idxJapanUSA = (Origin=="Japan"|Origin=="USA"); tbl = table(Model(idxJapanUSA),Origin(idxJapanUSA),MPG(idxJapanUSA),VariableNames=["Origin" "Model" "MPG"]);
Japan and the United States each manufacture a unique set of models. Therefore, the factor Model is nested in the factor Origin. Perform a two-way, nested ANOVA to test the null hypothesis that the car mileage is the same between the models and countries of origin.
aov = anova(tbl,"MPG ~ Origin + Model(Origin)")aov =
2-way anova, constrained (Type III) sums of squares.
MPG ~ 1 + Origin + Model(Origin)
SumOfSquares DF MeanSquares F pValue
____________ ___ ___________ ______ __________
Origin 18873 244 77.347 10.138 3.0582e-25
Model(Origin) 0 0 0 0 NaN
Error 633.26 83 7.6296
Total 19506 327
Properties, Methods
The small p-values indicate that the null hypothesis can be rejected at the 99% confidence level. Enough evidence exists to conclude that the model of the car and the country of origin have a statistically significant effect on the car mileage.
Algorithms
ANOVA partitions the total variation in the response data into two components:
Variation in the relationship between the factor data and the response data, as described by the ANOVA model. This variation is known as the sum of squares regression (SSR). The SSR is represented by the equation , where n is the number of observations in the sample, is the predicted value of observation i, and is the sample mean.
Variation in the data due to the ANOVA model error term, known as the sum of squares error (SSE). The SSE is represented by the equation , where is the value of observation i.
With the above partitioning, the total sum of squares (SST) is represented by
The anova function calculates the sum of
squares of a term () in the ANOVA model by measuring the reduction in the SSE
when the term is added to a comparison model. The comparison model is given by
aov.SumOfSquaresType (see SumOfSquaresType
for more information).
ANOVA uses SSE and to perform an F-test. For categorical main effects, the null hypothesis is that the term's coefficient is the same across all groups. For continuous and interaction terms, the null hypothesis is that the term's coefficient is zero. A zero coefficient means that the value of the term does not have an effect on the response data. The F-statistic is calculated as
In the above formula, is the degrees of freedom of a term, is the degrees of freedom of the error, and and are the mean squares of the term and error, respectively.
The anova function displays a component ANOVA table with rows
for the model terms and error. The columns of the ANOVA table are described as
follows:
| Column | Definition |
|---|---|
SumOfSquares | Sum of squares |
DF | Degrees of freedom |
MeanSquares | Mean squares, which is the ratio SumOfSquares/DF |
F | F-statistic, which is the source mean square to error mean square ratio |
pValue | p-value, which is the probability that the F-statistic, as computed under the null hypothesis, can take a value larger than the computed test-statistic value. anova derives this probability from the cdf of the F-distribution |
References
[1] Wackerly, D. D., W. Mendenhall, III, and R. L. Scheaffer. Mathematical Statistics with Applications, 7th ed. Belmont, CA: Brooks/Cole, 2008.
[2] Dunn, O. J., and V. A. Clark Hoboken. Applied Statistics: Analysis of Variance and Regression. NJ: John Wiley & Sons, Inc., 1974.
Version History
Introduced in R2022b
See Also
anova | anovan | anova2 | anova1 | N-Way ANOVA | One-Way ANOVA | Two-Way ANOVA
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.

Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list:
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)