Color-Based Segmentation Using K-Means Clustering
This example shows how to segment colors in an automated fashion using k-means clustering.
Clustering is a way to separate groups of objects. K-means clustering treats each object as having a location in space. It finds partitions such that objects within each cluster are as close to each other as possible, and as far from objects in other clusters as possible. You can use the imsegkmeans function to separate image pixels by value into clusters within a color space. This example performs k-means clustering of an image in the RGB and L*a*b* color spaces to show how using different color spaces can improve segmentation results.
Step 1: Read Image
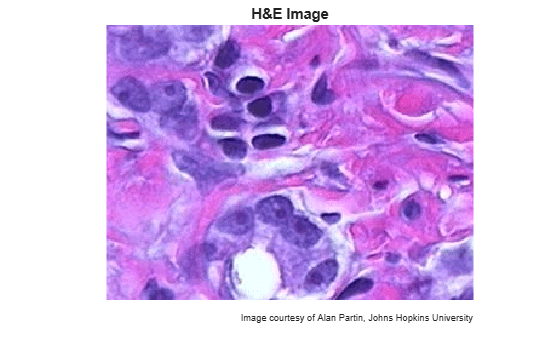
Read and display an image of tissue stained with hematoxylin and eosin (H&E). This staining method helps pathologists distinguish between tissue types that are stained blue-purple and pink.
he = imread("hestain.png"); imshow(he) title("H&E Image") text(size(he,2),size(he,1)+15, ... "Image courtesy of Alan Partin, Johns Hopkins University", ... FontSize=7,HorizontalAlignment="right")
Step 2: Classify Colors in RBG Color Space Using K-Means Clustering
Segment the image into three regions using k-means clustering in the RGB color space. For each pixel in the input image, the imsegkmeans function returns a label corresponding to a cluster.
Display the label image as an overlay on the original image. The label image incorrectly groups white, light blue-purple, and light pink regions together. Because the RGB color space combines brightness and color information within each channel (red, green, blue), lighter versions of two different colors are closer together and more challenging to segment than darker versions of the same two colors.
numColors = 3;
L = imsegkmeans(he,numColors);
B = labeloverlay(he,L);
imshow(B)
title("Labeled Image RGB")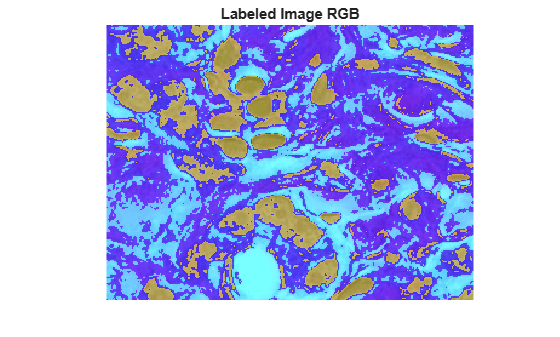
Step 3: Convert Image from RGB Color Space to L*a*b* Color Space
The L*a*b* color space separates image luminosity and color. This makes it easier to segment regions by color, independent of lightness. The color space is also more consistent with human visual perception of the distinct white, blue-purple, and pink regions in the image.
The L*a*b* color space is derived from the CIE XYZ tristimulus values. The L*a*b* space consists of the luminosity layer L*, the chromaticity layer a* that indicates where a color falls along the red-green axis, and the chromaticity layer b* that indicates where a color falls along the blue-yellow axis. All of the color information is in the a* and b* layers.
Convert the image to the L*a*b* color space by using the rgb2lab function.
lab_he = rgb2lab(he);
Step 4: Classify Colors in a*b* Space Using K-Means Clustering
To segment the image using only color information, limit the image to the a* and b* values in lab_he. Convert the image to data type single for use with the imsegkmeans function. Use the imsegkmeans function to separate the image pixels into three clusters. Set the value of the NumAttempts name-value argument to repeat clustering three times with different initial cluster centroid positions to avoid fitting to a local minimum.
ab = lab_he(:,:,2:3); ab = im2single(ab); pixel_labels = imsegkmeans(ab,numColors,NumAttempts=3);
Display the label image as an overlay on the original image. The new label image more clearly separates the white, blue-purple, and pink stained tissue regions.
B2 = labeloverlay(he,pixel_labels);
imshow(B2)
title("Labeled Image a*b*")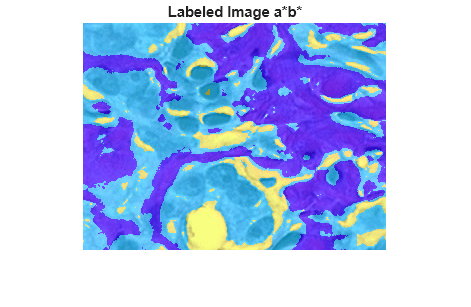
Step 5: Create Images That Segment H&E Image by Color
Using pixel_labels, you can separate objects in the original image hestain.png by color, resulting in three masked images.
mask1 = pixel_labels == 1;
cluster1 = he.*uint8(mask1);
imshow(cluster1)
title("Objects in Cluster 1");
mask2 = pixel_labels == 2;
cluster2 = he.*uint8(mask2);
imshow(cluster2)
title("Objects in Cluster 2");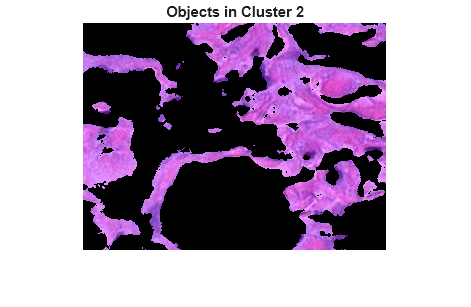
mask3 = pixel_labels == 3;
cluster3 = he.*uint8(mask3);
imshow(cluster3)
title("Objects in Cluster 3");
Step 6: Segment Nuclei
Cluster 3 contains only the blue objects. Notice that there are dark and light blue objects. You can separate dark blue from light blue using the L* layer in the L*a*b* color space. The cell nuclei are dark blue.
The L* layer contains the brightness value of each pixel. Extract the brightness values of the pixels in this cluster and threshold them with a global threshold by using the imbinarize function. The mask idx_light_blue gives the indices of light blue pixels.
L = lab_he(:,:,1); L_blue = L.*double(mask3); L_blue = rescale(L_blue); idx_light_blue = imbinarize(nonzeros(L_blue));
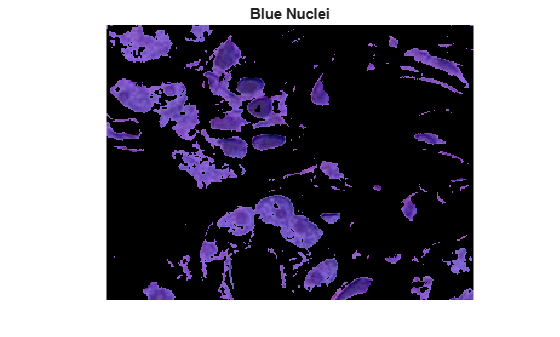
Copy the mask of blue objects, mask3, and then remove the light blue pixels from the mask. Apply the new mask to the original image and display the result. Only dark blue cell nuclei are visible.
blue_idx = find(mask3);
mask_dark_blue = mask3;
mask_dark_blue(blue_idx(idx_light_blue)) = 0;
blue_nuclei = he.*uint8(mask_dark_blue);
imshow(blue_nuclei)
title("Blue Nuclei")