Process Blocked Images Efficiently Using Mask
This example shows how to process a blocked image efficiently by using a mask to isolate regions of interest (ROIs).
Some sources of large images have meaningful data in only a small portion of the image. You can improve total processing time by limiting processing to the ROI containing meaningful data. Use a mask to define ROIs. A mask is a logical image in which true pixels represent the ROI.
In the blocked image workflow, the mask represents the same spatial region as the image data, but it does not need to be the same size as the image. To further improve the efficiency of the workflow, create a mask from a coarse image, especially one that fits in memory. Then, use the coarse mask to process the finer images.
Create a blocked image using a modified version of image tumor_091.tif from the CAMELYON16 data set. The original image is a training image of a lymph node containing tumor tissue. The original image has eight resolution levels, and the finest level has resolution 53760-by-61440. The modified image has only three coarse resolution levels. The spatial referencing of the modified image has been adjusted to enforce a consistent aspect ratio and to register features at each level.
bim = blockedImage("tumor_091R.tif");Display the blocked image by using the imageshow function.
imageshow(bim);
Create Mask
Determine the image size at the coarsest level. The coarsest level is the last level in the blocked image.
coarseLevel = bim.NumLevels; coarseLevelSize = bim.Size(coarseLevel,:)
coarseLevelSize = 1×3
625 670 3
Get the image at the coarsest resolution level.
imLowRes = gather(bim);
You can generate a mask from the coarse image using the Image Segmenter app. Because the app expects a grayscale input image, you must extract the lightness channel from the coarse image.
imLowResL = rgb2lightness(imLowRes);
To run the Image Segmenter app, enter this command in the Command Window:
imageSegmenter(imLowResL)
After you define the mask, export the mask as BW, or export the code that the app uses to create the mask. This section of the example uses code exported from the app. Run this code to create and display a mask from the coarse input image.
%---------------------------------------------------- % Normalize input data to range in [0,1]. Xmin = min(imLowResL(:)); Xmax = max(imLowResL(:)); if isequal(Xmax,Xmin) imLowResL = 0*imLowResL; else imLowResL = (imLowResL - Xmin) ./ (Xmax - Xmin); end % Threshold image - global threshold BW = imbinarize(imLowResL); % Invert mask BW = imcomplement(BW); % Open mask with square width = 3; se = strel("square", width); BW = imopen(BW, se); %---------------------------------------------------- imageshow(BW)
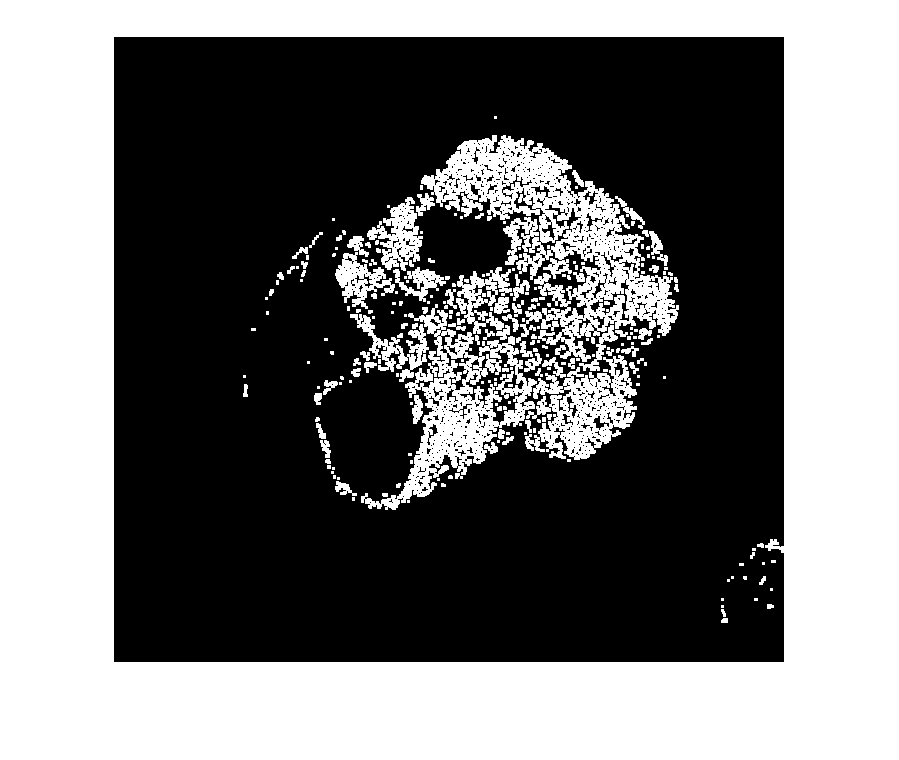

Create a blocked image from the mask with the same spatial referencing as the input mask.
bmask = blockedImage(BW,WorldEnd=bim.WorldEnd(3,1:2));
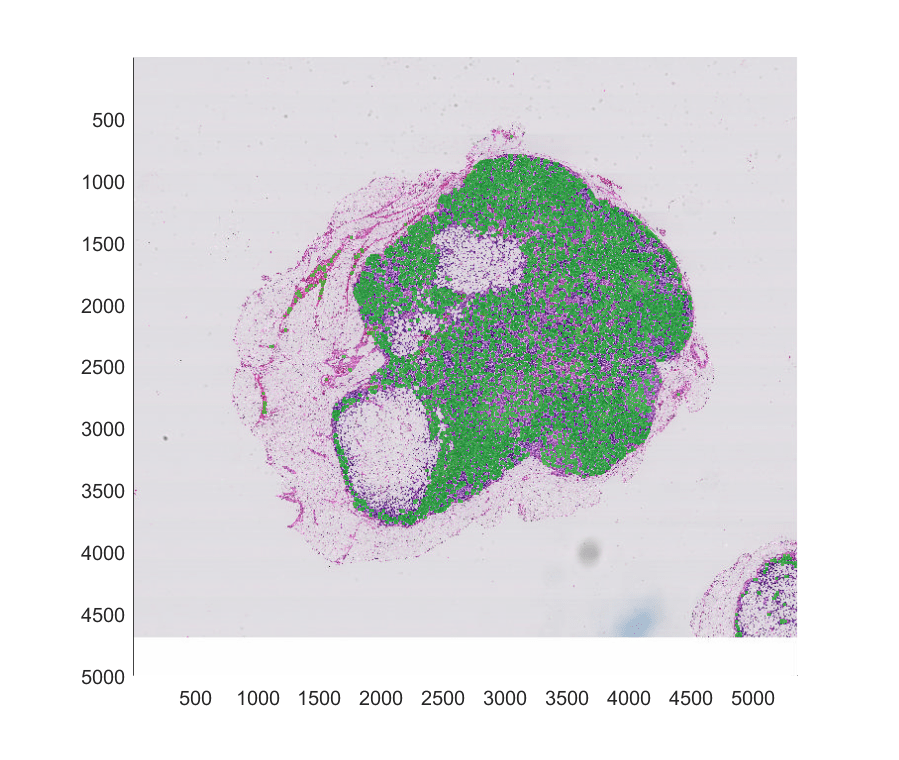
Display the mask as a translucent green overlay on the original blocked image.
imageshow(bim,OverlayData=bmask,OverlayDisplayRangeMode="data-range", ... OverlayAlphamap = [0 0.5],OverlayColormap=[0 1 0]);

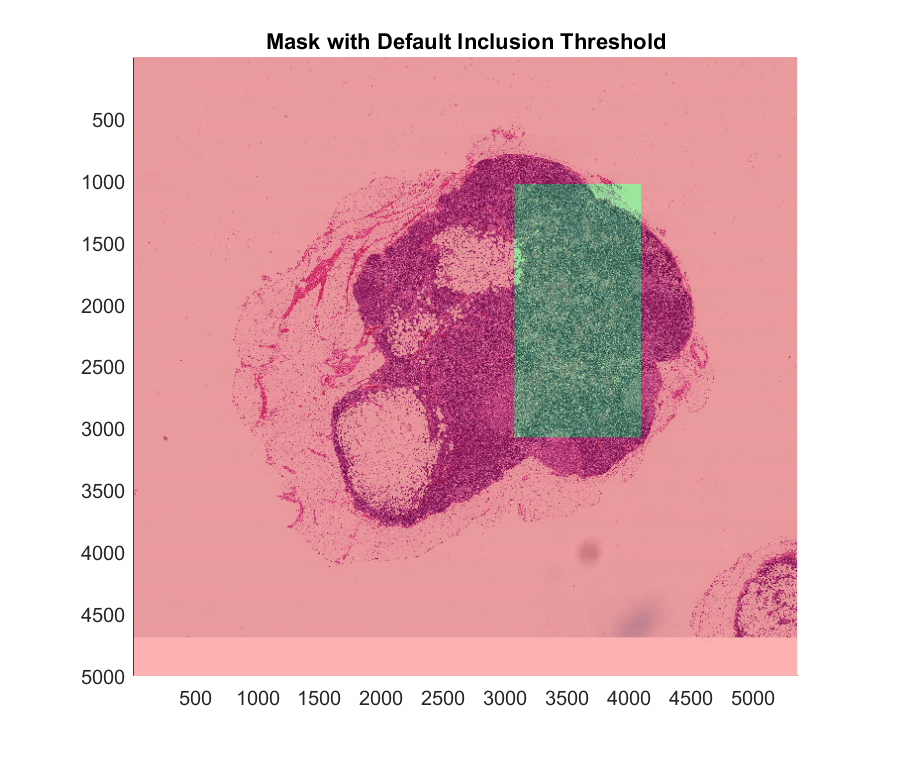
Adjust Inclusion Threshold to Cover Region of Interest
The apply function processes blocked images one block at a time. You can use the InclusionThreshold name-value argument with the mask to specify which blocks the apply function uses. The inclusion threshold specifies the percentage of mask pixels that must be true for apply to process the block. To process more blocks of the image, decrease the inclusion threshold. You can also process all blocks that have at least a single true pixel in the mask. To use this option, specify the InclusionThreshold name-value argument as 0. Note that not all blocks of the image are included.
Using the mask with any value of InclusionThreshold decreases the total execution time because apply processes only a subset of blocks from the full image. The benefit of using a mask is more significant at higher resolutions and as the processing pipeline increases in complexity.
Measure the execution time of filtering the full image.
tic bout = apply(bim,@(bs)imnlmfilt(bs.Data,DegreeOfSmoothing=15)); tFullProcessing = toc;
Measure the execution time of filtering only the blocks within the ROI.
bls = selectBlockLocations(bim,Mask=bmask,InclusionThreshold=0); tic boutMasked = apply(bim, ... @(bs)imnlmfilt(bs.Data,DegreeOfSmoothing=15), ... BlockLocationSet=bls); tMaskedProcessing = toc; boutMasked = makeMultiLevel2D(boutMasked,Scales=[1 0.5 0.3 0.1]); hbim = imageshow(boutMasked); defaultBlockSize = bim.BlockSize(1,:); hbim.Parent.Title = "Processed Image Using Mask with Default BlockSize = [" + ... defaultBlockSize + "]";
Compare the execution time of processing the full image to the execution time of processing only the blocks in the ROI.
disp("Speedup using mask: " + ... num2str(tFullProcessing/tMaskedProcessing) + "x");
Speedup using mask: 2.274x
Adjust Block Size to Follow Contours of Region of Interest
You can decrease the block size to create a tighter wrap around the ROI. For some block sizes, this reduces the execution time because apply processes fewer pixels outside the ROI. However, if the block size is too small, then performance decreases because the overhead of processing a larger number of blocks offsets the reduction in the number of pixels processed.
To specify the block size, use the BlockSize name-value argument. Measure the execution time of filtering all the blocks within the ROI with a decreased block size.
blockSize = [512 512]; bls = selectBlockLocations(bim,Mask=bmask,InclusionThreshold=0,BlockSize=blockSize); tic boutMasked = apply(bim, ... @(bs)imnlmfilt(bs.Data,DegreeOfSmoothing=15), ... BlockLocationSet=bls); tSmallerBlockProcessing = toc; boutMasked = makeMultiLevel2D(boutMasked,Scales=[1 0.5 0.3 0.1]); hbim = imageshow(boutMasked); hbim.Parent.Title = "Processed Image Using Mask with BlockSize = [" + ... blockSize + "]";

Compare the execution time of processing the entire ROI with smaller blocks to the execution time of processing the entire ROI with the original blocks.
disp("Additional speedup using mask with decreased block size: " + ... num2str(tMaskedProcessing/tSmallerBlockProcessing) + "x");
Additional speedup using mask with decreased block size: 1.2767x
See Also
blockedImage | bigimageshow | apply