pwelch
Welch’s power spectral density estimate
Syntax
Description
[
returns Welch's PSD estimate of the signal pxx,f] = pwelch(x,win,nOverlap,freqSpec)x and the
frequencies f (rad/sample), where the
pwelch function:
Uses
winto divide the signal into segments and windows them.Uses the overlap length specified in
nOverlapto overlap samples between adjoining segments.Computes the discrete Fourier transform (DFT) of each windowed segment over the number of DFT points or at the frequencies specified in
freqSpec.
To use default values for any of these input arguments, specify
them as empty, [].
[___] = pwelch(___,
also specifies the frequency range, spectrum type, and trace mode for any of the
previous syntaxes. You can specify any combination of these input
arguments.freqRange,spectrumType,trace)
pwelch(___) with no output arguments plots
the Welch PSD estimate or power spectrum in the current figure window.
Examples
Obtain the Welch PSD estimate of an input signal consisting of a discrete-time sinusoid with an angular frequency of rad/sample with additive white noise.
Create a sine wave with an angular frequency of rad/sample with additive white noise. Reset the random number generator for reproducible results. The signal has a length samples.
rng("default")
n = 0:319;
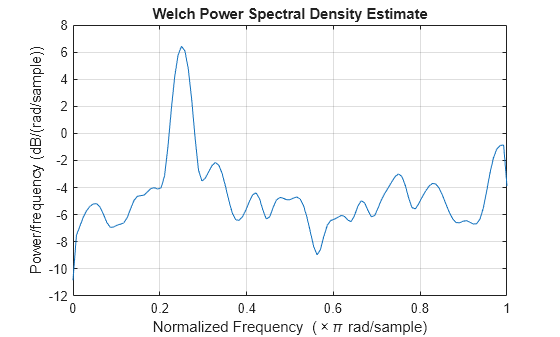
x = cos(pi/4*n) + randn(size(n));Obtain the Welch PSD estimate using the default Hamming window and DFT length. The default segment length is 71 samples and the DFT length is the 256 points yielding a frequency resolution of rad/sample. Because the signal is real-valued, the periodogram is one-sided and there are 256/2+1 points. Plot the Welch PSD estimate.
pxx = pwelch(x); pwelch(x)
Repeat the computation.
Divide the signal into sections of length . This action is equivalent to dividing the signal into the longest possible segments to obtain as close to but not exceed 8 segments with 50% overlap.
Window the sections using a Hamming window.
Specify 50% overlap between contiguous sections
To compute the FFT, use points, where .
Verify that the two approaches give identical results.
Nx = length(x); nsc = floor(Nx/4.5); nov = floor(nsc/2); nff = max(256,2^nextpow2(nsc)); pxxt = pwelch(x,hamming(nsc),nov,nff); maxerr = max(abs(abs(pxxt(:)) - abs(pxx(:))))
maxerr = 0
Divide the signal into 8 sections of equal length, with 50% overlap between sections. Specify the same FFT length as in the preceding step. Compute the Welch PSD estimate and verify that it gives the same result as the previous two procedures.
ns = 8; ov = 0.5; lsc = floor(Nx/(ns-(ns-1)*ov)); pxxt8 = pwelch(x,lsc,floor(ov*lsc),nff); maxerr8 = max(abs(abs(pxxt8(:)) - abs(pxx(:))))
maxerr8 = 0
Obtain the Welch PSD estimate of an input signal consisting of a discrete-time sinusoid with an angular frequency of rad/sample with additive white noise.
Create a sine wave with an angular frequency of rad/sample with additive white noise. Reset the random number generator for reproducible results. The signal has 512 samples.
rng("default")
n = 0:511;
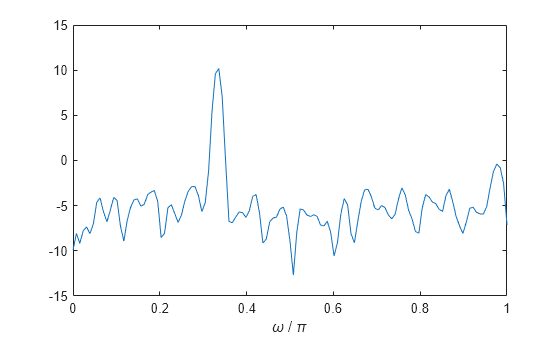
x = cos(pi/3*n) + randn(size(n));Obtain the Welch PSD estimate dividing the signal into segments 132 samples in length. The signal segments are multiplied by a Hamming window 132 samples in length. The number of overlapped samples is not specified, so it is set to 132/2 = 66. The DFT length is 256 points, yielding a frequency resolution of rad/sample. Because the signal is real-valued, the PSD estimate is one-sided and there are 256/2+1 = 129 points. Plot the PSD as a function of normalized frequency.
segmentLength = 132; [pxx,w] = pwelch(x,segmentLength); plot(w/pi,pow2db(pxx)) xlabel("Normalized Frequency (\times \pi rad/sample)") title("Welch Power Spectral Density Estimate")
Obtain the Welch PSD estimate of an input signal consisting of a discrete-time sinusoid with an angular frequency of rad/sample with additive white noise.
Create a sine wave with an angular frequency of rad/sample with additive white noise. Reset the random number generator for reproducible results. The signal is 320 samples in length.
rng("default")
n = 0:319;
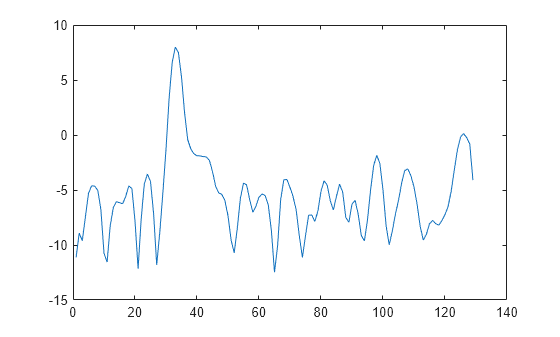
x = cos(pi/4*n) + randn(size(n));Obtain the Welch PSD estimate dividing the signal into segments 100 samples in length. The signal segments are multiplied by a Hamming window 100 samples in length. The number of overlapped samples is 25. The DFT length is 256 points yielding a frequency resolution of rad/sample. Because the signal is real-valued, the PSD estimate is one-sided and there are 256/2+1 points.
segmentLength = 100; noverlap = 25; pxx = pwelch(x,segmentLength,noverlap); plot(pow2db(pxx)) xlabel("Frequency Samples") title("Welch Power Spectral Density Estimate")
Obtain the Welch PSD estimate of an input signal consisting of a discrete-time sinusoid with an angular frequency of rad/sample with additive white noise.
Create a sine wave with an angular frequency of rad/sample with additive white noise. Reset the random number generator for reproducible results. The signal is 320 samples in length.
rng("default")
n = 0:319;
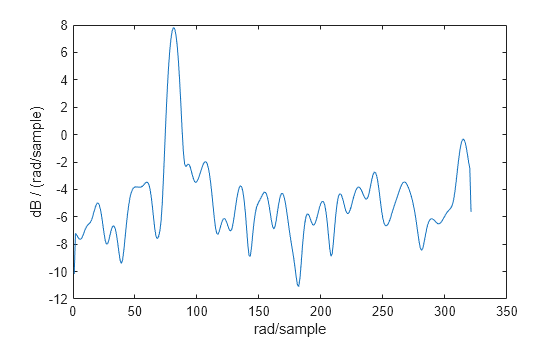
x = cos(pi/4*n) + randn(size(n));Obtain the Welch PSD estimate dividing the signal into segments 100 samples in length. Use the default overlap of 50%. Specify the DFT length to be 640 points so that the frequency of rad/sample corresponds to a DFT bin (bin 81). Because the signal is real-valued, the PSD estimate is one-sided and there are 640/2+1 points.
segmentLength = 100; nfft = 640; pxx = pwelch(x,segmentLength,[],nfft); plot(pow2db(pxx)) xlabel("Frequency Points") title("Welch Power Spectral Density Estimate")
Create a signal consisting of a 100 Hz sinusoid in additive N(0,1) white noise. Reset the random number generator for reproducible results. The sample rate is 1 kHz and the signal is 5 seconds in duration.
rng("default")
Fs = 1000;
t = 0:1/Fs:5-1/Fs;
x = cos(2*pi*100*t) + randn(size(t));Obtain Welch's overlapped segment averaging PSD estimate of the preceding signal. Use a segment length of 500 samples with 300 overlapped samples. Use 500 DFT points so that 100 Hz falls directly on a DFT bin. Input the sample rate to output a vector of frequencies in Hz. Plot the result.
[pxx,f] = pwelch(x,500,300,500,Fs); plot(f,pow2db(pxx)) xlabel("Frequency (Hz)") ylabel("PSD (dB/Hz)")

Create a signal consisting of three noisy sinusoids and a chirp, sampled at 200 kHz for 0.1 second. The frequencies of the sinusoids are 1 kHz, 10 kHz, and 20 kHz. The sinusoids have different amplitudes and noise levels. The noiseless chirp has a frequency that starts at 20 kHz and increases linearly to 30 kHz during the sampling.
rng("default")
Fs = 200e3;
Fc = [1 10 20]'*1e3;
Ns = 0.1*Fs;
t = (0:Ns-1)/Fs;
x = [1 1/10 10]*sin(2*pi*Fc*t) + [1/200 1/2000 1/20]*randn(3,Ns);
x = x + chirp(t,20e3,t(end),30e3);Compute the Welch PSD estimate and the maximum-hold and minimum-hold spectra of the signal. Plot the results.
[pxx,f] = pwelch(x,[],[],[],Fs); pmax = pwelch(x,[],[],[],Fs,"maxhold"); pmin = pwelch(x,[],[],[],Fs,"minhold"); plot(f,pow2db(pxx)) hold on plot(f,pow2db([pmax pmin]),":") hold off xlabel("Frequency (Hz)") ylabel("PSD (dB/Hz)") legend(["pwelch" "maxhold" "minhold"])
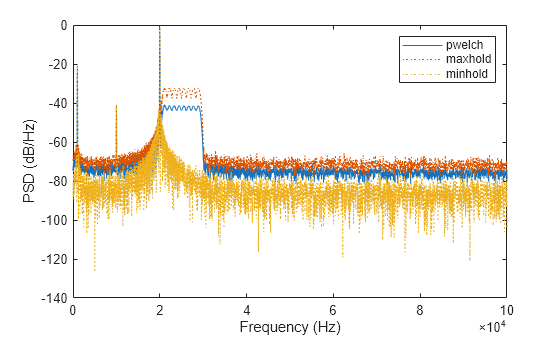
Repeat the procedure, this time computing centered power spectrum estimates.
[pxx,f] = pwelch(x,[],[],[],Fs,"centered","power"); pmax = pwelch(x,[],[],[],Fs,"maxhold","centered","power"); pmin = pwelch(x,[],[],[],Fs,"minhold","centered","power"); plot(f,pow2db(pxx)) hold on plot(f,pow2db([pmax pmin]),":") hold off xlabel("Frequency (Hz)") ylabel("Power (dB)") legend(["pwelch" "maxhold" "minhold"])
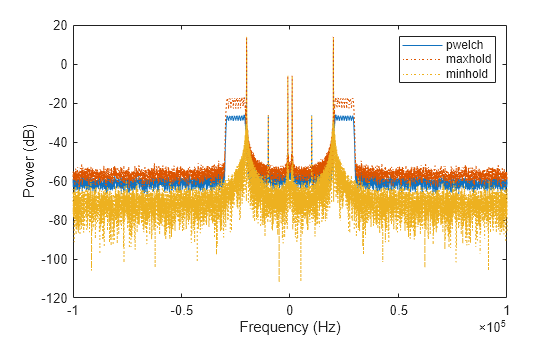
This example illustrates the use of confidence bounds with Welch's overlapped segment averaging (WOSA) PSD estimate. While not a necessary condition for statistical significance, frequencies in Welch's estimate where the lower confidence bound exceeds the upper confidence bound for surrounding PSD estimates clearly indicate significant oscillations in the time series.
Create a signal consisting of the superposition of 100 Hz and 150 Hz sine waves in additive white N(0,1) noise. The amplitude of the two sine waves is 1. The sample rate is 1 kHz. Reset the random number generator for reproducible results.
rng("default")
Fs = 1000;
t = 0:1/Fs:1-1/Fs;
x = cos(2*pi*100*t) + sin(2*pi*150*t) + randn(size(t));Obtain the WOSA estimate with 95%-confidence bounds. Set the segment length equal to 200 and overlap the segments by 50% (100 samples).
L = 200; win = hamming(L); nOverlap = 100; [pxx,f,pxxc] = pwelch(x,win,nOverlap,200,Fs,ConfidenceLevel=0.95);
Plot the WOSA PSD estimate along with the confidence interval and zoom in on the frequency region of interest near 100 and 150 Hz. The lower confidence bound in the immediate vicinity of 100 and 150 Hz is significantly above the upper confidence bound outside the vicinity of 100 and 150 Hz.
plot(f,pow2db(pxx)) hold on plot(f,pow2db(pxxc),"-.",Color=[0.866 0.329 0]) hold off xlim([25 250]) xlabel("Frequency (Hz)") ylabel("PSD (dB/Hz)") title("Welch Estimate with 95%-Confidence Bounds")
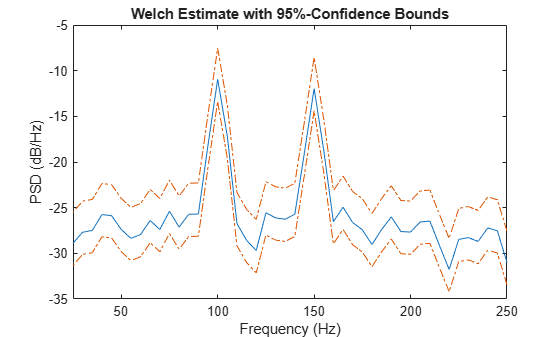
Create a signal consisting of a 100 Hz sinusoid in additive white noise. Reset the random number generator for reproducible results. The sample rate is 1 kHz and the signal is 5 seconds in duration.
rng("default")
Fs = 1000;
t = 0:1/Fs:5-1/Fs;
noisevar = 1/4;
x = cos(2*pi*100*t) + sqrt(noisevar)*randn(size(t));Obtain the DC-centered power spectrum using Welch's method. Use a segment length of 500 samples with 300 overlapped samples and a DFT length of 500 points.
[pxx,f] = pwelch(x,500,300,500,Fs,"centered","power");
Plot the result. The power at -100 and 100 Hz is close to the expected power of 1/4 for a real-valued sine wave with an amplitude of 1. The deviation from 1/4 is due to the effect of the additive noise.
plot(f,pow2db(pxx)) xlabel("Frequency (Hz)") ylabel("Magnitude (dB)") grid
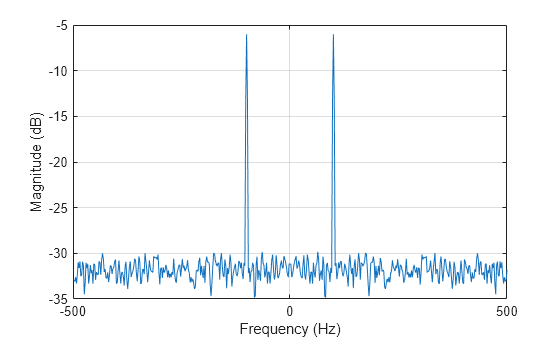
Generate 1024 samples of a multichannel signal consisting of three sinusoids in additive white Gaussian noise. The sinusoids' frequencies are , , and rad/sample. Estimate the PSD of the signal using Welch's method and plot it.
N = 1024;
n = 0:N-1;
w = pi./[2;3;4];
rng("default")
x = cos(w*n)' + randn(length(n),3);
pwelch(x)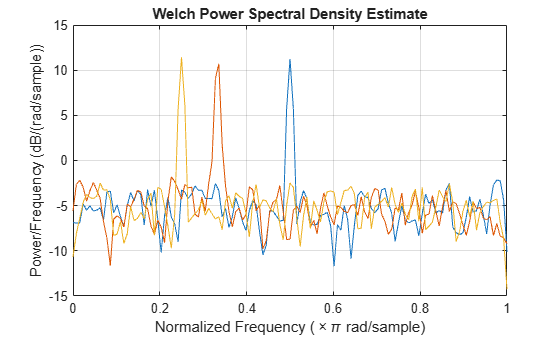
Since R2026a
Plot the Welch power spectral density (PSD) estimate and Welch power spectrum for four signals in the specified target axes and panel containers.
Create four oscillating signals with a sample rate of 10 kHz for three seconds.
Fs = 10e3;
t = 0:1/Fs:3;
x1 = sinc(Fs/2.5*(t-mean(t)));
x2 = sum(cos(2*pi*600*[1 3 5 7]'.*t),1) + randn(size(t))/1e4;
x3 = exp(1j*pi*sin(4*t)*Fs/10);
x4 = chirp(t,Fs/10,t(end),Fs/2.5,"quadratic");Plot Welch PSD Estimate and Power Spectrum in Target Axes
Create two axes in the southwestern and northeastern corners of a new figure window.
fig = figure; ax1 = axes(fig,Position=[0.09 0.1 0.52 0.45]); ax2 = axes(fig,Position=[0.55 0.7 0.42 0.25]);
Plot the Welch PSD estimate and Welch power spectrum of the signals x1 and x2 in the southwestern and northeastern axes of the figure, respectively. Use a 256-sample Kaiser window, an overlap length of 220 samples, and 512 DFT points.
g = kaiser(256,5);
ol = 220;
nfft = 512;
pwelch(x1,g,ol,nfft,Fs,Parent=ax1)
pwelch(x2,g,ol,nfft,Fs,"power",Parent=ax2)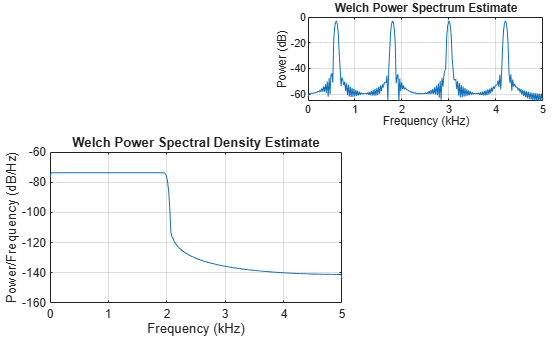
Plot Welch PSD Estimate in Target UI Axes
Create an axes in the northwestern corner of a new UI figure window.
uif = uifigure(Position=[100 100 720 540]); ax3 = uiaxes(uif,Position=[5 305 300 200]);
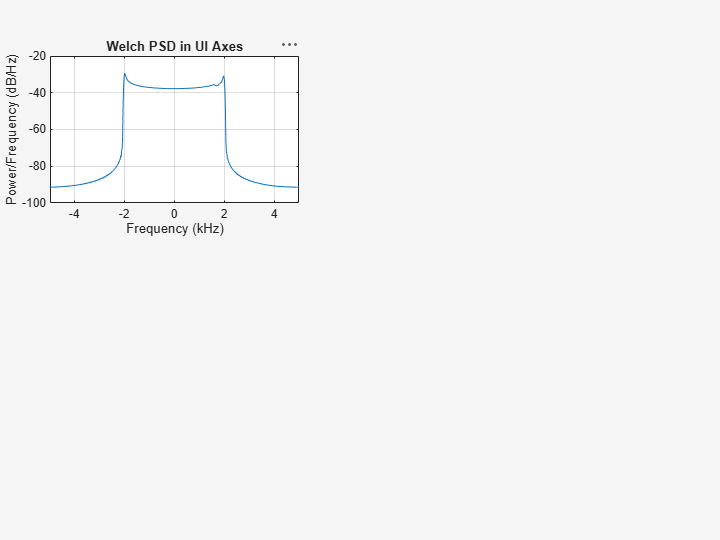
Plot the Welch PSD estimate of the signal x3 on the figure axes. Display the frequencies centered at 0 kHz.
pwelch(x3,g,ol,nfft,Fs,"centered",Parent=ax3) title(ax3,"Welch PSD in UI Axes")
Plot Welch PSD Estimate in Target Panel Container
Add a panel container in the southeastern corner of the UI figure window.
p = uipanel(uif,Position=[300 5 400 325], ... Title="Welch PSD in Panel Container", ... BackgroundColor="white");
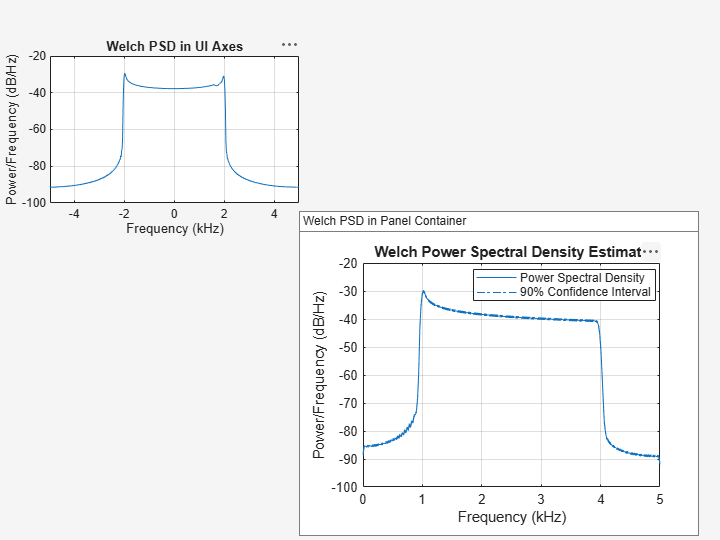
Plot the Welch PSD estimate of the signal x4 on the panel container. Set the confidence level to 90%.
pwelch(x4,g,ol,nfft,Fs,ConfidenceLevel=0.9,Parent=p)
Input Arguments
Input signal, specified as a vector or as a matrix. If you specify x as a
matrix, then pwelch treats its columns as independent
channels.
Example: cos(pi/4*(0:159))+randn(1,160) is
a single-channel row-vector signal.
Example: cos(pi./[4;2]*(0:159))'+randn(160,2) is
a two-channel signal.
Data Types: single | double
Complex Number Support: Yes
Window, specified as empty ([]), a positive integer, or a
vector.
The pwelch function divides the input signal
x into segments and uses a window to compute the
element-by-element product of the segment samples and the window values. The window
values depend on the how you specify this argument.
Empty (
[]) —pwelchuses a Hamming window of length such thatxis divided into eight segments withnOverlapoverlapping samples.Positive integer —
pwelchdividesxinto segments of lengthwinand uses a Hamming window of that length.Vector —
pwelchdividesxinto segments of the same length as the vectorwinand uses the window specified inwin.
If the length of x cannot be divided exactly into an
integer number of segments with nOverlap overlapping samples, then
pwelch truncates x
accordingly.
For a list of available windows, see Windows.
Note
The default Hamming window has a 42.5 dB sidelobe attenuation, which may mask spectral content below this value (relative to the peak spectral content). Choosing different windows enables you to make tradeoffs between resolution (for example, using a rectangular window) and sidelobe attenuation (for example, using a Hann window). See Window Designer for more details.
Example: hann(N+1) and (1-cos(2*pi*(0:N)'/N))/2
both specify a Hann window of length N + 1.
Data Types: single | double
Number of overlapped samples, specified as empty ([])
or a nonnegative integer.
Empty (
[]) —pwelchuses a number that produces 50% overlap between segments.If the segment length is unspecified, the function sets
nOverlapto ⌊Nx / 4.5⌋, where Nx is the length of the input signal and the ⌊ ⌋ symbols denote the floor function. This action is equivalent to dividing the signal into the longest possible segments to obtain as close to but not exceed eight segments with 50% overlap.Nonnegative integer —
pwelchoverlaps adjoining segments using the number of samples specified in this argument.If
winis scalar, thennOverlapmust be smaller thanwin.If
winis a vector, thennOverlapmust be smaller than the length ofwin.
Frequency specification, specified as one of these values:
Empty,
[]—pwelchcalculates the spectral estimates using a number of DFT points equal to the greater of 256 or the next power of two that is equal or larger than the window length.Positive integer —
pwelchcalculates the spectral estimates using the number of DFT points specified in this argument.If
freqSpecis greater than the segment length, thenpwelchpads the segment data with zeros.If
freqSpecis less than the segment length, thenpwelchpartitions the segment modulofreqSpecand sums the resulting segment frames.
Vector of at least two elements —
pwelchcalculates the spectral estimates at the frequencies specified infreqSpec.If you specify the sample rate,
Fs,pwelchassumesfreqSpecas cyclical frequencies in the same units as ofFs.If you do not specify
Fs,pwelchassumesfreqSpecas normalized frequencies in rad/sample.
Example: freqSpec = 512 specifies 512 DFT points.
Example: freqSpec = pi./[8 4 2] specifies a vector of three normalized frequencies.
Data Types: single | double
Sample rate, specified as a positive scalar. The sample rate is the number of samples per unit time. If the unit of time is seconds, then the sample rate is in Hz.
If you specify
Fsas empty[], thenpwelchassumes that the input signalxhas a sample rate of 1 Hz.If you do not specify
Fs, thenpwelchassumes that the input signalxhas a sample rate of 2π rad/sample.
Frequency range of the PSD estimate or power spectrum, specified as
"onesided", "twosided", or
"centered".
The pwelch function returns
pxx with a number of rows and frequency interval
that depends on the value specified in freqRange,
whether the number of DFT points freqSpec is even or
odd, and whether Fs is specified or not.
freqRange | freqSpec | Number of Rows in
pxx | Frequency Interval for
| |
|---|---|---|---|---|
|
| |||
"onesided"(default if x is real-valued) | Even | freqSpec/2 + 1 | [0,π] rad/sample | [0,Fs/2] cycles/unit
time |
| Odd | (freqSpec + 1)/2 | [0,π) rad/sample | [0,Fs/2) cycles/unit
time | |
"twosided"(default if x is complex-valued) | Even or odd | freqSpec | [0,2π) rad/sample | [0,Fs) cycles/unit time |
"centered" | Even | freqSpec | (–π,π] rad/sample | (–Fs/2,Fs/2]
cycles/unit time |
| Odd | (–π,π) rad/sample | (–Fs/2,Fs/2)
cycles/unit time | ||
Note
This argument is not supported if you specify
freqSpecas a vector of cyclical or normalized frequencies.If you specify the
"onesided"value,pwelchmultiplies the power by 2 at all frequencies except 0 and the Nyquist frequency to conserve the total power.The
"onesided"value is not supported ifxis complex-valued.
Data Types: char | string
Power spectrum scaling, specified as one of these values:
"psd"—pwelchreturns the power spectral density."power"—pwelchscales each estimate of the PSD by the equivalent noise bandwidth of the window and returns an estimate of the power at each frequency.
This table shows the scaling relation between the PSD
estimate and power spectrum estimate, either of both returned in
pxx, given an input signal x,
window vector win, overlap length
nOverlap, frequency specification
freqSpec, and sample rate
Fs.
| Sample Rate | Scaling Relation |
|---|---|
Fs specified | psd = pwelch(x,win,nOverlap,freqSpec,Fs,"psd"); pow = pwelch(x,win,nOverlap,freqSpec,Fs,"power"); pow is equivalent to
psd*enbw(win,Fs).
win is a window
vector. |
Fs unspecified | psd = pwelch(x,win,nOverlap,freqSpec,"psd"); pow = pwelch(x,win,nOverlap,freqSpec,"power"); pow is equivalent to
psd*enbw(win,2*pi). |
Trace mode, specified as one of these values:
"mean"—pwelchreturns Welch's spectrum estimate of each input channel by averaging the power spectrum estimates across all the segments at each frequency bin."maxhold"—pwelchreturns the maximum-hold spectrum of each input channel by selecting the maximum power spectrum value across segments at each frequency bin."minhold"—pwelchreturns the minimum-hold spectrum of each input channel by selecting the minimum power spectrum value across segments at each frequency bin.
Coverage probability for the PSD estimate, specified as a scalar in the range (0,1).
If you specify ConfidenceLevel=p and pxxc,
the function outputs pxxc with the lower and upper bounds of the
p × 100% interval estimate for the true
PSD.
Since R2026a
Target parent container, specified as an Axes object, a
UIAxes object,
or a Panel
object.
If you specify Parent=h, the
pwelch function plots the Welch PSD estimate
or power spectrum on the specified target parent container, whether you call
the function with or without output arguments.
For more information about target containers and the parent-child
relationship in MATLAB® graphics, see Graphics Object Hierarchy. For
more information about using Parent in
UIAxes and Panel objects to design
apps, see Plot Spectral Representations of Signal in App Designer.
Example: h = axes(figure,Position=[0.1 0.1 0.6 0.5])
defines an axes parent container. When you specify
pwelch(x,[],[],[],Parent=h), the function plots
the Welch PSD of the input signal x in the parent
container h.
Output Arguments
PSD estimate or power spectrum, returned as a real-valued, nonnegative column vector or matrix.
Each column of
pxxis the PSD estimate or the or power spectrum of the corresponding column ofxand depending on how you specifyspectrumType.The units of the PSD estimate are in squared-magnitude units of the input signal per unit frequency. For example, if the input data
xis in volts, you specify the sample rateFsin hertz, and you assume a resistance of 1 Ω, then the PSD estimate is in watts per hertz.The units of the power spectrum are in squared-magnitude units of the input signal. For example, if the input data
xis in volts and you assume a resistance of 1 Ω, then the PSD estimate is in watts.
Frequencies associated with the PSD estimate, returned as a real-valued column vector.
If you specify
Fs, thenfcontains cyclical frequencies in Hz.If you do not specify
Fs, thenfcontains normalized frequencies in rad/sample.
Confidence bounds, returned as a real-valued matrix.
pxxchas the same number of rows as inpxx.pxxchas twice as many columns aspxx.Odd-numbered columns contain the lower bounds of the confidence intervals.
Even-numbered columns contain the upper bounds of the confidence intervals.
Thus,
pxxc(m,2*n-1)is the lower confidence bound andpxxc(m,2*n)is the upper confidence bound corresponding to the estimatepxx(m,n).The coverage probability of the confidence intervals is determined by the value of the
pinput.
Data Types: single | double
More About
The periodogram is not a consistent estimator of the true power spectral density of a wide-sense stationary process. Welch’s technique to reduce the variance of the periodogram breaks the input signal into overlapping segments.
Welch’s method computes a modified periodogram for each segment and then averages these estimates to produce the estimate of the power spectral density. Because the process is wide-sense stationary and Welch’s method uses PSD estimates of different segments of the input signal, the modified periodograms represent approximately uncorrelated estimates of the true PSD and averaging reduces the variability.
The segments are typically multiplied by a window function, such as a Hamming window, so that Welch’s method amounts to averaging modified periodograms. Because the segments usually overlap, data values at the beginning and end of the segment tapered by the window in one segment, occur away from the ends of adjacent segments. This guards against the loss of information caused by windowing.
References
[1] Hayes, Monson H. Statistical Digital Signal Processing and Modeling. New York: John Wiley & Sons, 1996.
[2] Stoica, Petre, and Randolph Moses. Spectral Analysis of Signals. Upper Saddle River, NJ: Prentice Hall, 2005.
Extended Capabilities
The
pwelch function supports tall arrays with the following usage
notes and limitations:
The input
xmust not be a tall row vectorThe
winargument must always be specified.
For more information, see Tall Arrays.
C/C++ Code Generation
Generate C and C++ code using MATLAB® Coder™.
GPU Code Generation
Generate CUDA® code for NVIDIA® GPUs using GPU Coder™.
The pwelch function supports
thread-based environments with these usage notes and limitations:
The syntax with no output arguments is not supported.
For more information, see Run MATLAB Functions in Thread-Based Environment.
This function fully supports GPU arrays. For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced before R2006aThe pwelch function supports plotting output in a parent
axes or panel container specified using the Parent input
argument.
The pwelch function supports code generation for
graphical processing units (GPUs). You must have MATLAB
Coder™ and GPU Coder™ to generate CUDA® code.
The pwelch function supports single-precision
variable-size window inputs for code generation.
The pwelch function supports single-precision
inputs.
You can now use the Create
Plot Live Editor task to visualize the output of
pwelch interactively. You can select different chart
types and set optional parameters. The task also automatically generates code that
becomes part of your live script.
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)