CalinskiHarabaszEvaluation
Calinski-Harabasz criterion clustering evaluation object
Description
CalinskiHarabaszEvaluation is an object consisting of sample data
(X), clustering data (OptimalY), and Calinski-Harabasz
criterion values (CriterionValues) used to
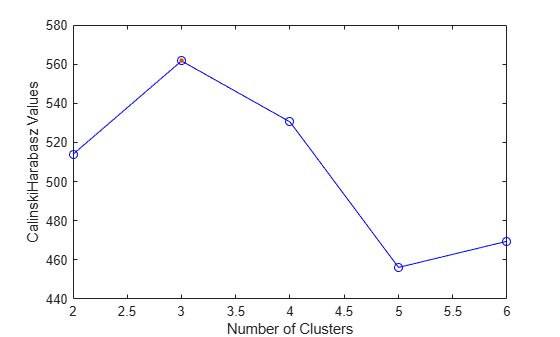
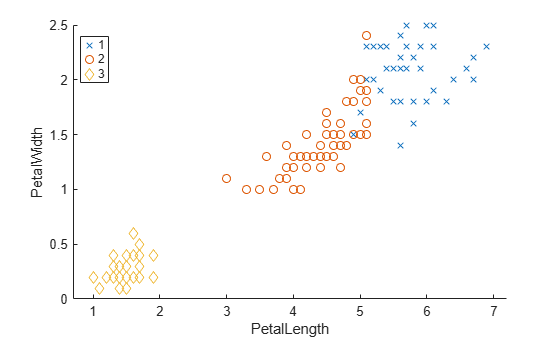
evaluate the optimal number of clusters (OptimalK). The Calinski-Harabasz
criterion is sometimes called the variance ratio criterion (VRC). Well-defined clusters have a
large between-cluster variance and a small within-cluster variance. The optimal number of
clusters corresponds to the solution with the highest Calinski-Harabasz index value. For more
information, see Calinski-Harabasz Criterion.
Creation
Create a Calinski-Harabasz criterion clustering evaluation object by using the evalclusters function and specifying the criterion as
"CalinskiHarabasz".
You can then use compact to create a compact version of the
Calinski-Harabasz criterion clustering evaluation object. The function removes the contents of
the properties X, OptimalY, and
Missing.
Properties
Object Functions
Examples
More About
References
[1] Calinski, T., and J. Harabasz. “A dendrite method for cluster analysis.” Communications in Statistics. Vol. 3, No. 1, 1974, pp. 1–27.
Version History
Introduced in R2013b
See Also
evalclusters | DaviesBouldinEvaluation | GapEvaluation | SilhouetteEvaluation