survival
Calculate survival of Cox proportional hazards model
Syntax
Description
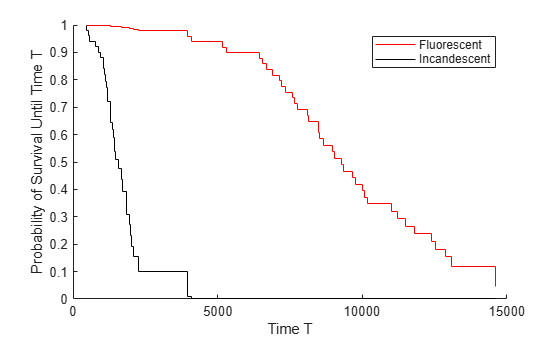
s = survival(coxMdl)coxMdl. The survival function at time t is the
estimated probability of survival until time t. The term
baseline refers to the survival function at the determined baseline
of the predictors. This value is stored in coxMdl.Baseline, and the
default value is the mean of the data set used for training.
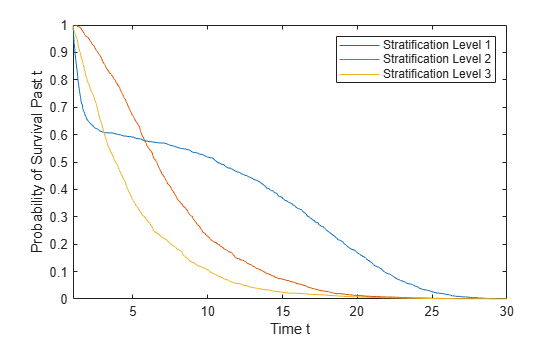
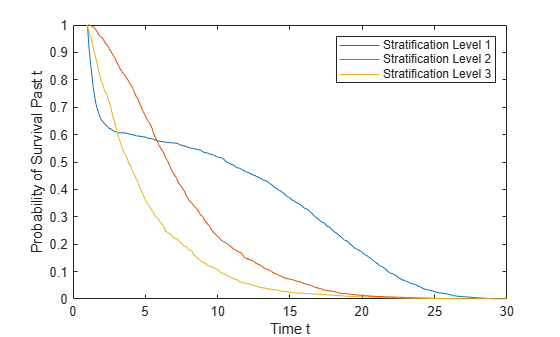
s = survival(coxMdl,X,Stratification)Stratification. You must have one row in
Stratification for each row in X.
Note
When you train coxMdl using stratification variables and
pass predictor variables X,
survival also requires you to pass
stratification variables.
s = survival(___,Name,Value)survival(CoxMdl,"Time",T) computes the survival at times
T.
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Version History
Introduced in R2021a