crossval
Estimate loss using cross-validation
Syntax
Description
err = crossval(criterion,X,y,'Predfun',predfun)predfun based on the specified criterion,
either 'mse' (mean squared error) or 'mcr'
(misclassification rate). The rows of X and y
correspond to observations, and the columns of X correspond to
predictor variables.
For more information, see General Cross-Validation Steps for predfun.
values = crossval(fun,X)fun, applied to
the data in X. The rows of X correspond to
observations, and the columns of X correspond to variables.
For more information, see General Cross-Validation Steps for fun.
___ = crossval(___,
specifies cross-validation options using one or more name-value pair arguments in addition
to any of the input argument combinations and output arguments in previous syntaxes. For
example, Name,Value)'KFold',5 specifies to perform 5-fold cross-validation.
Examples
Compute the mean squared error of a regression model by using 10-fold cross-validation.
Load the carsmall data set. Put the acceleration, horsepower, weight, and miles per gallon (MPG) values into the matrix data. Remove any rows that contain NaN values.
load carsmall
data = [Acceleration Horsepower Weight MPG];
data(any(isnan(data),2),:) = [];Specify the last column of data, which corresponds to MPG, as the response variable y. Specify the other columns of data as the predictor data X. Add a column of ones to X when your regression function uses regress, as in this example.
Note: regress is useful when you simply need the coefficient estimates or residuals of a regression model. If you need to investigate a fitted regression model further, create a linear regression model object by using fitlm. For an example that uses fitlm and crossval, see Compute Mean Absolute Error Using Cross-Validation.
y = data(:,4); X = [ones(length(y),1) data(:,1:3)];
Create the custom function regf (shown at the end of this example). This function fits a regression model to training data and then computes predicted values on a test set.
Note: If you use the live script file for this example, the regf function is already included at the end of the file. Otherwise, you need to create this function at the end of your .m file or add it as a file on the MATLAB® path.
Compute the default 10-fold cross-validation mean squared error for the regression model with predictor data X and response variable y.
rng('default') % For reproducibility cvMSE = crossval('mse',X,y,'Predfun',@regf)
cvMSE = 17.5399
This code creates the function regf.
function yfit = regf(Xtrain,ytrain,Xtest) b = regress(ytrain,Xtrain); yfit = Xtest*b; end
Compute the misclassification error of a logistic regression model trained on numeric and categorical predictor data by using 10-fold cross-validation.
Load the patients data set. Specify the numeric variables Diastolic and Systolic and the categorical variable Gender as predictors, and specify Smoker as the response variable.
load patients
X1 = Diastolic;
X2 = categorical(Gender);
X3 = Systolic;
y = Smoker;Create the custom function classf (shown at the end of this example). This function fits a logistic regression model to training data and then classifies test data.
Note: If you use the live script file for this example, the classf function is already included at the end of the file. Otherwise, you need to create this function at the end of your .m file or add it as a file on the MATLAB® path.
Compute the 10-fold cross-validation misclassification error for the model with predictor data X1, X2, and X3 and response variable y. Specify 'Stratify',y to ensure that training and test sets have roughly the same proportion of smokers.
rng('default') % For reproducibility err = crossval('mcr',X1,X2,X3,y,'Predfun',@classf,'Stratify',y)
err = 0.1100
This code creates the function classf.
function pred = classf(X1train,X2train,X3train,ytrain,X1test,X2test,X3test) Xtrain = table(X1train,X2train,X3train,ytrain, ... 'VariableNames',{'Diastolic','Gender','Systolic','Smoker'}); Xtest = table(X1test,X2test,X3test, ... 'VariableNames',{'Diastolic','Gender','Systolic'}); modelspec = 'Smoker ~ Diastolic + Gender + Systolic'; mdl = fitglm(Xtrain,modelspec,'Distribution','binomial'); yfit = predict(mdl,Xtest); pred = (yfit > 0.5); end
For a given number of clusters, compute the cross-validated sum of squared distances between observations and their nearest cluster center. Compare the results for one through ten clusters.
Load the fisheriris data set. X is the matrix meas, which contains flower measurements for 150 different flowers.
load fisheriris
X = meas;Create the custom function clustf (shown at the end of this example). This function performs the following steps:
Standardize the training data.
Separate the training data into
kclusters.Transform the test data using the training data mean and standard deviation.
Compute the distance from each test data point to the nearest cluster center, or centroid.
Compute the sum of the squares of the distances.
Note: If you use the live script file for this example, the clustf function is already included at the end of the file. Otherwise, you need to create the function at the end of your .m file or add it as a file on the MATLAB® path.
Create a for loop that specifies the number of clusters k for each iteration. For each fixed number of clusters, pass the corresponding clustf function to crossval. Because crossval performs 10-fold cross-validation by default, the software computes 10 sums of squared distances, one for each partition of training and test data. Take the sum of those values; the result is the cross-validated sum of squared distances for the given number of clusters.
rng('default') % For reproducibility cvdist = zeros(5,1); for k = 1:10 fun = @(Xtrain,Xtest)clustf(Xtrain,Xtest,k); distances = crossval(fun,X); cvdist(k) = sum(distances); end
Plot the cross-validated sum of squared distances for each number of clusters.
plot(cvdist) xlabel('Number of Clusters') ylabel('CV Sum of Squared Distances')
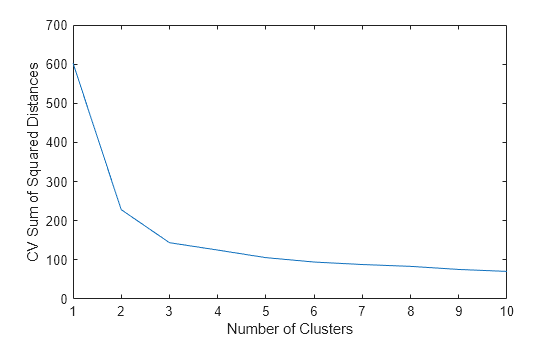
In general, when determining how many clusters to use, consider the greatest number of clusters that corresponds to a significant decrease in the cross-validated sum of squared distances. For this example, using two or three clusters seems appropriate, but using more than three clusters does not.
This code creates the function clustf.
function distances = clustf(Xtrain,Xtest,k) [Ztrain,Zmean,Zstd] = zscore(Xtrain); [~,C] = kmeans(Ztrain,k); % Creates k clusters Ztest = (Xtest-Zmean)./Zstd; d = pdist2(C,Ztest,'euclidean','Smallest',1); distances = sum(d.^2); end
Compute the mean absolute error of a regression model by using 10-fold cross-validation.
Load the carsmall data set. Specify the Acceleration and Displacement variables as predictors and the Weight variable as the response.
load carsmall
X1 = Acceleration;
X2 = Displacement;
y = Weight;Create the custom function regf (shown at the end of this example). This function fits a regression model to training data and then computes predicted car weights on a test set. The function compares the predicted car weight values to the true values, and then computes the mean absolute error (MAE) and the MAE adjusted to the range of the test set car weights.
Note: If you use the live script file for this example, the regf function is already included at the end of the file. Otherwise, you need to create this function at the end of your .m file or add it as a file on the MATLAB® path.
By default, crossval performs 10-fold cross-validation. For each of the 10 training and test set partitions of the data in X1, X2, and y, compute the MAE and adjusted MAE values using the regf function. Find the mean MAE and mean adjusted MAE.
rng('default') % For reproducibility values = crossval(@regf,X1,X2,y)
values = 10×2
319.2261 0.1132
342.3722 0.1240
214.3735 0.0902
174.7247 0.1128
189.4835 0.0832
249.4359 0.1003
194.4210 0.0845
348.7437 0.1700
283.1761 0.1187
210.7444 0.1325
mean(values)
ans = 1×2
252.6701 0.1129
This code creates the function regf.
function errors = regf(X1train,X2train,ytrain,X1test,X2test,ytest) tbltrain = table(X1train,X2train,ytrain, ... 'VariableNames',{'Acceleration','Displacement','Weight'}); tbltest = table(X1test,X2test,ytest, ... 'VariableNames',{'Acceleration','Displacement','Weight'}); mdl = fitlm(tbltrain,'Weight ~ Acceleration + Displacement'); yfit = predict(mdl,tbltest); MAE = mean(abs(yfit-tbltest.Weight)); adjMAE = MAE/range(tbltest.Weight); errors = [MAE adjMAE]; end
Compute the misclassification error of a classification tree by using principal component analysis (PCA) and 5-fold cross-validation.
Load the fisheriris data set. The meas matrix contains flower measurements for 150 different flowers. The species variable lists the species for each flower.
load fisheririsCreate the custom function classf (shown at the end of this example). This function fits a classification tree to training data and then classifies test data. Use PCA inside the function to reduce the number of predictors used to create the tree model.
Note: If you use the live script file for this example, the classf function is already included at the end of the file. Otherwise, you need to create this function at the end of your .m file or add it as a file on the MATLAB® path.
Create a cvpartition object for stratified 5-fold cross-validation. By default, cvpartition ensures that training and test sets have roughly the same proportions of flower species.
rng('default') % For reproducibility cvp = cvpartition(species,'KFold',5);
Compute the 5-fold cross-validation misclassification error for the classification tree with predictor data meas and response variable species.
cvError = crossval('mcr',meas,species,'Predfun',@classf,'Partition',cvp)
cvError = 0.1067
This code creates the function classf.
function yfit = classf(Xtrain,ytrain,Xtest) % Standardize the training predictor data. Then, find the % principal components for the standardized training predictor % data. [Ztrain,Zmean,Zstd] = zscore(Xtrain); [coeff,scoreTrain,~,~,explained,mu] = pca(Ztrain); % Find the lowest number of principal components that account % for at least 95% of the variability. n = find(cumsum(explained)>=95,1); % Find the n principal component scores for the standardized % training predictor data. Train a classification tree model % using only these scores. scoreTrain95 = scoreTrain(:,1:n); mdl = fitctree(scoreTrain95,ytrain); % Find the n principal component scores for the transformed % test data. Classify the test data. Ztest = (Xtest-Zmean)./Zstd; scoreTest95 = (Ztest-mu)*coeff(:,1:n); yfit = predict(mdl,scoreTest95); end
Create a confusion matrix from the 10-fold cross-validation results of a discriminant analysis model.
Note: Use classify when training speed is a concern. Otherwise, use fitcdiscr to create a discriminant analysis model. For an example that shows the same workflow as this example, but uses fitcdiscr, see Create Confusion Matrix Using Cross-Validation Predictions.
Load the fisheriris data set. X contains flower measurements for 150 different flowers, and y lists the species for each flower. Create a variable order that specifies the order of the flower species.
load fisheriris
X = meas;
y = species;
order = unique(y)order = 3×1 cell
{'setosa' }
{'versicolor'}
{'virginica' }
Create a function handle named func for a function that completes the following steps:
Take in training data (
Xtrainandytrain) and test data (Xtestandytest).Use the training data to create a discriminant analysis model that classifies new data (
Xtest). Create this model and classify new data by using theclassifyfunction.Compare the true test data classes (
ytest) to the predicted test data values, and create a confusion matrix of the results by using theconfusionmatfunction. Specify the class order by using'Order',order.
func = @(Xtrain,ytrain,Xtest,ytest)confusionmat(ytest, ... classify(Xtest,Xtrain,ytrain),'Order',order);
Create a cvpartition object for stratified 10-fold cross-validation. By default, cvpartition ensures that training and test sets have roughly the same proportions of flower species.
rng('default') % For reproducibility cvp = cvpartition(y,'Kfold',10);
Compute the 10 test set confusion matrices for each partition of the predictor data X and response variable y. Each row of confMat corresponds to the confusion matrix results for one test set. Aggregate the results and create the final confusion matrix cvMat.
confMat = crossval(func,X,y,'Partition',cvp);
cvMat = reshape(sum(confMat),3,3)cvMat = 3×3
50 0 0
0 48 2
0 1 49
Plot the confusion matrix as a confusion matrix chart by using confusionchart.
confusionchart(cvMat,order)
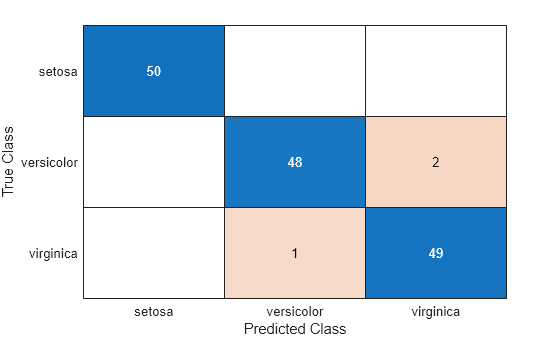
Input Arguments
Type of error estimate, specified as either 'mse' or
'mcr'.
| Value | Description |
|---|---|
'mse' | Mean squared error (MSE) — Appropriate for regression algorithms only |
'mcr' | Misclassification rate, or proportion of misclassified observations — Appropriate for classification algorithms only |
Data set, specified as a column vector, matrix, or array. The rows of
X correspond to observations, and the columns of
X generally correspond to variables. If you pass multiple data
sets X1,...,XN to crossval, then all data sets
must have the same number of rows.
Data Types: single | double | logical | char | string | cell | categorical
Response data, specified as a column vector or character array. The rows of
y correspond to observations, and y must
have the same number of rows as the predictor data X or
X1,...,XN.
Data Types: single | double | logical | char | string | cell | categorical
Prediction function, specified as a function handle. You must create this function as an anonymous function, a function defined at the end of the .m or .mlx file containing the rest of your code, or a file on the MATLAB® path.
This table describes the required function syntax, given the type of predictor data
passed to crossval.
| Value | Predictor Data | Function Syntax |
|---|---|---|
@myfunction | X |
function yfit = myfunction(Xtrain,ytrain,Xtest) % Calculate predicted response ... end
|
@myfunction | X1,...,XN |
function yfit = myfunction(X1train,...,XNtrain,ytrain,X1test,...,XNtest) % Calculate predicted response ... end
|
Example: @(Xtrain,ytrain,Xtest)(Xtest*regress(ytrain,Xtrain));
Data Types: function_handle
Function to cross-validate, specified as a function handle. You must create this function as an anonymous function, a function defined at the end of the .m or .mlx file containing the rest of your code, or a file on the MATLAB path.
This table describes the required function syntax, given the type of data passed to
crossval.
| Value | Data | Function Syntax |
|---|---|---|
@myfunction | X |
function value = myfunction(Xtrain,Xtest) % Calculation of value ... end
|
@myfunction | X1,...,XN |
function value = myfunction(X1train,...,XNtrain,X1test,...,XNtest) % Calculation of value ... end
|
Data Types: function_handle
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: crossval('mcr',meas,species,'Predfun',@classf,'KFold',5,'Stratify',species)
specifies to compute the stratified 5-fold cross-validation misclassification rate for the
classf function with predictor data meas and
response variable species.
Fraction or number of observations used for holdout validation, specified as the
comma-separated pair consisting of 'Holdout' and a scalar value in
the range (0,1) or a positive integer scalar.
If the
Holdoutvaluepis a scalar in the range (0,1), thencrossvalrandomly selects and reserves approximatelyp*100% of the observations as test data.If the
Holdoutvaluepis a positive integer scalar, thencrossvalrandomly selects and reservespobservations as test data.
In either case, crossval then trains the model
specified by either fun or predfun using the
rest of the data. Finally, the function uses the test data along with the trained
model to compute either values or
err.
You can use only one of these four name-value pair arguments:
Holdout, KFold,
Leaveout, and Partition.
Example: 'Holdout',0.3
Example: 'Holdout',50
Data Types: single | double
Number of folds for k-fold cross-validation, specified as the comma-separated pair
consisting of 'KFold' and a positive integer scalar greater than
1.
If you specify 'KFold',k, then crossval
randomly partitions the data into k sets. For each set, the
function reserves the set as test data, and trains the model specified by either
fun or predfun using the other
k – 1 sets. crossval then uses the test data
along with the trained model to compute either values or
err.
You can use only one of these four name-value pair arguments:
Holdout, KFold,
Leaveout, and Partition.
Example: 'KFold',5
Data Types: single | double
Leave-one-out cross-validation, specified as the comma-separated pair consisting
of 'Leaveout' and 1.
If you specify 'Leaveout',1, then for each observation,
crossval reserves the observation as test data, and trains the
model specified by either fun or predfun
using the other observations. The function then uses the test observation along with
the trained model to compute either values or
err.
You can use only one of these four name-value pair arguments:
Holdout, KFold,
Leaveout, and Partition.
Example: 'Leaveout',1
Data Types: single | double
Number of Monte Carlo repetitions for validation, specified as the comma-separated
pair consisting of 'MCReps' and a positive integer scalar. If the
first input of crossval is 'mse' or
'mcr' (see criterion), then
crossval returns the mean MSE or misclassification rate across
all Monte Carlo repetitions. Otherwise, crossval concatenates the
values from all Monte Carlo repetitions along the first dimension.
If you specify both Partition and
MCReps, then the first Monte Carlo repetition uses the partition
information in the cvpartition object, and the software calls the
repartition object function to generate
new partitions for each of the remaining Monte Carlo repetitions.
If the Leaveout value is 1, the
Partition value is a cvpartition object of
type 'leaveout' or 'resubstitution', or the
Partition value is a custom cvpartition object
(that is, the IsCustom property is set to 1),
then the software sets the MCReps value to
1.
Example: 'MCReps',5
Data Types: single | double
Cross-validation partition, specified as the comma-separated pair consisting of
'Partition' and a cvpartition partition object
created by cvpartition. The partition object
specifies the type of cross-validation and the indexing for the training and test
sets.
When you use crossval, you cannot specify both
Partition and Stratify. Instead, directly
specify a stratified partition when you create the cvpartition
partition object.
You can use only one of these four name-value pair arguments:
Holdout, KFold,
Leaveout, and Partition.
Variable specifying the groups used for stratification, specified as the
comma-separated pair consisting of 'Stratify' and a column vector
with the same number of rows as the data X or
X1,...,XN.
When you specify Stratify, both the training and test sets
have roughly the same class proportions as in the Stratify
vector. The software treats NaNs, empty character vectors, empty
strings, <missing> values, and <undefined>
values in Stratify as missing data values, and ignores the
corresponding rows of the data.
A good practice is to use stratification when you use cross-validation with classification algorithms. Otherwise, some test sets might not include observations for all classes.
When you use crossval, you cannot specify both
Partition and Stratify. Instead, directly
specify a stratified partition when you create the cvpartition
partition object.
Data Types: single | double | logical | string | cell | categorical
Options for computing in parallel and setting random streams, specified as a
structure. Create the Options structure using statset. This table lists the option fields and their
values.
| Field Name | Value | Default |
|---|---|---|
UseParallel | Set this value to true to run computations in
parallel. | false |
UseSubstreams | Set this value to To compute
reproducibly, set | false |
Streams | Specify this value as a RandStream object or
cell array of such objects. Use a single object except when the
UseParallel value is true
and the UseSubstreams value is
false. In that case, use a cell array that
has the same size as the parallel pool. | If you do not specify Streams, then
crossval uses the default stream or
streams. |
Note
You need Parallel Computing Toolbox™ to run computations in parallel.
Example: Options=statset(UseParallel=true,UseSubstreams=true,Streams=RandStream("mlfg6331_64"))
Data Types: struct
Output Arguments
Mean squared error or misclassification rate, returned as a numeric scalar. The type
of error depends on the criterion value.
Loss values, returned as a column vector or matrix. Each row of
values corresponds to the output of fun for
one partition of training and test data.
If the output returned by fun is multidimensional, then
crossval reshapes the output and fits it into one row of
values. For an example, see Create Confusion Matrix Using Cross-Validation.
Tips
A good practice is to use stratification (see
Stratify) when you use cross-validation with classification algorithms. Otherwise, some test sets might not include observations for all classes.
Algorithms
When you use predfun, the crossval function
typically performs 10-fold cross-validation as follows:
Split the observations in the predictor data
Xand the response variableyinto 10 groups, each of which has approximately the same number of observations.Use the last nine groups of observations to train a model as specified in
predfun. Use the first group of observations as test data, pass the test predictor data to the trained model, and compute predicted values as specified inpredfun. Compute the error specified bycriterion.Use the first group and the last eight groups of observations to train a model as specified in
predfun. Use the second group of observations as test data, pass the test data to the trained model, and compute predicted values as specified inpredfun. Compute the error specified bycriterion.Proceed in a similar manner until each group of observations is used as test data exactly once.
Return the mean error estimate as the scalar
err.
When you use fun, the crossval function
typically performs 10-fold cross-validation as follows:
Split the data in
Xinto 10 groups, each of which has approximately the same number of observations.Use the last nine groups of data to train a model as specified in
fun. Use the first group of data as a test set, pass the test set to the trained model, and compute some value (for example, loss) as specified infun.Use the first group and the last eight groups of data to train a model as specified in
fun. Use the second group of data as a test set, pass the test set to the trained model, and compute some value as specified infun.Proceed in a similar manner until each group of data is used as a test set exactly once.
Return the 10 computed values as the vector
values.
Alternative Functionality
Many classification and regression functions allow you to perform cross-validation directly.
When you use fit functions such as
fitcsvm,fitctree, andfitrtree, you can specify cross-validation options by using name-value pair arguments. Alternatively, you can first create models with these fit functions and then create a partitioned object by using thecrossvalobject function. Use thekfoldLossandkfoldPredictobject functions to compute the loss and predicted values for the partitioned object. For more information, seeClassificationPartitionedModelandRegressionPartitionedModel.You can also specify cross-validation options when you perform lasso or elastic net regularization using
lassoandlassoglm.
Extended Capabilities
To run in parallel, specify the Options name-value argument in the call to
this function and set the UseParallel field of the
options structure to true using
statset:
Options=statset(UseParallel=true)
For more information about parallel computing, see Run MATLAB Functions with Automatic Parallel Support (Parallel Computing Toolbox).
Version History
Introduced in R2008a
See Also
cvpartition | pca | regress | classify | kmeans | confusionmat
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)