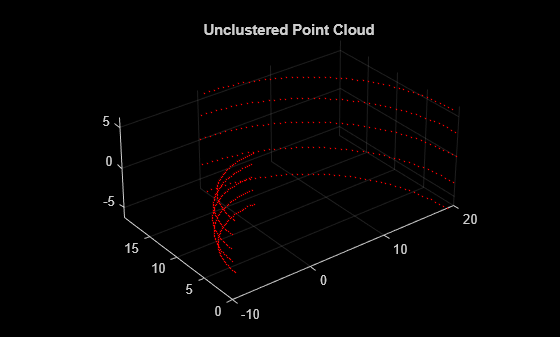
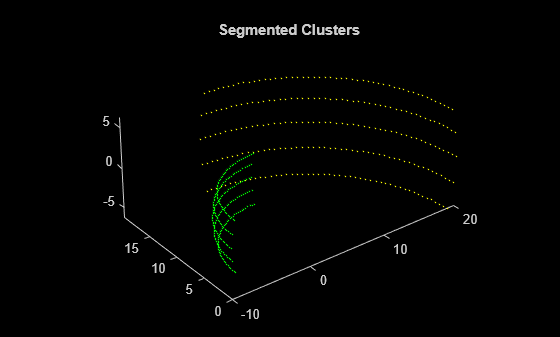
segmentLidarData
Segment organized 3-D range data into clusters
Syntax
Description
labels = segmentLidarData(ptCloud,distThreshold)ptCloud into clusters. The
function assigns an integer cluster label to each point in the point cloud and
returns the cluster label of all points in labels.
The function groups two neighboring points into the same cluster if their
Euclidean distance is less than distThreshold or if the angle
between the sensor and two neighboring points is at least 5 degrees.
labels = segmentLidarData(ptCloud,distThreshold,angleThreshold)angleThreshold.
[
also returns the number of clusters.labels,numClusters] = segmentLidarData(___)
[___] = segmentLidarData(___,NumClusterPoints=[1,Inf])
also sets the minimum and maximum number of points in each cluster, specified as a
2-element vector or as a scalar value. When you specify
NumClusterPoints as a scalar, the maximum number of points in
the cluster is unrestricted. The function sets the labels to
0 when clusters are outside of the specified range.
Examples
Input Arguments
Output Arguments
Algorithms
The segmentLidarData function uses distance and angle thresholds to
cluster neighboring points. The function groups two neighboring points into the same
cluster if their Euclidean distance is less than the input
distThreshold or if the angle between the sensor and
neighboring points is greater than or equal to the input
angleThreshold. If you do not specify
angleThreshold, the function sets this angle to
5 degrees.
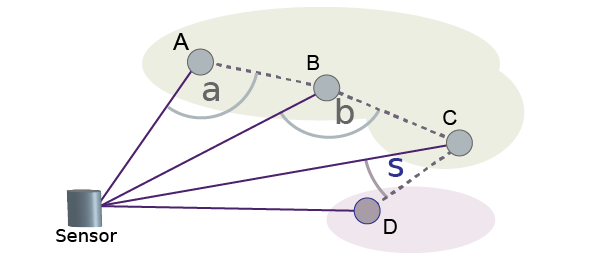
For example, suppose angleThreshold is set to
90. Because angles α and β in the figure are both greater than
the specified threshold of 90 degrees, the function groups points A, B, and C into the
same cluster. Because angle σ is less than the 90-degree threshold, the function groups
point D into a separate cluster. Each angle the function uses for clustering is formed
by the line from a point to the sensor and the line from that same point to the
neighboring point
References
[1] Bogoslavskyi, I. “Efficient Online Segmentation for Sparse 3D Laser Scans.” Journal of Photogrammetry, Remote Sensing and Geoinformation Science. Vol. 85, Issue 1, 2017, pp. 41–52.
Extended Capabilities
Version History
Introduced in R2018a
See Also
pointCloud | pcsegdist | pcfitplane | velodyneFileReader | segmentGroundFromLidarData