You are now following this Submission
- You will see updates in your followed content feed
- You may receive emails, depending on your communication preferences
normxcorr2_general computes the normalized cross-correlation of matrices TEMPLATE and A. The resulting matrix C contains correlation coefficients and its values may range from -1.0 to 1.0.
Limitations of normxcorr2:
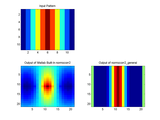
The documentation of normxcorr2 states that, "The matrix A must be larger than the matrix TEMPLATE for the normalization to be meaningful." It is implemented following the details of the paper "Fast Normalized Cross-Correlation", by J. P. Lewis, Industrial Light & Magic. This approach assumes the template is small relative to the image and proceeds to calculate the normalization across the entire template. This leads to correct computations wherever the template is wholly overlapping with the image, but the computation is incorrect in the borders of the output (the border size is proportional to the template size). This problem is therefore worse for larger templates to the point that, when the template is the same size as the image, the only correct value is at the center pixel (where the images are fully overlapping). Thus, if normxcorr2 is used for such things as registering images of the same size, the result will be incorrect.
The new normxcorr2_general:
normxcorr2_general is more general than normxcorr2 in that it gives correct results everywhere regardless of the relative size of A and TEMPLATE. It accomplishes this by computing the normalized correlation only in the overlap regions between the two matrices. Thus, the result is correct for all locations of correlation. The result is the same as if the NCC were carried out in the spatial domain (which would take a long time to compute for large matrices).
Cite As
Dirk Padfield (2026). Generalized Normalized Cross Correlation (https://www.mathworks.com/matlabcentral/fileexchange/29005-generalized-normalized-cross-correlation), MATLAB Central File Exchange. Retrieved .
General Information
- Version 1.3.0.0 (5.3 KB)
MATLAB Release Compatibility
- Compatible with any release
Platform Compatibility
- Windows
- macOS
- Linux
| Version | Published | Release Notes | Action |
|---|---|---|---|
| 1.3.0.0 | Made the code independent of the image processing toolbox (IPT). Note that the IPT is needed to run the example since normxcorr2 is part of that toolbox. However, normxcorr2_general does not require the IPT. |
||
| 1.2.0.0 | Added a new optional parameter. requiredNumberOfOverlapPixels sets to 0 all locations in C computed from positions where A and T overlap less than requiredNumberOfOverlapPixels. |
||
| 1.0.0.0 |