You are now following this Submission
- You will see updates in your followed content feed
- You may receive emails, depending on your communication preferences
First thing to do:
Run a tutorial code, tutorial.m
Documentation:
http://2000.jukuin.keio.ac.jp/shimazaki/res/kernel.html
see also sskernel for optimization of a fixed kernel bandwidth and sshist for histogram optimization.
% [y,t,optw,gs,C,confb95,yb] = ssvkernel(x,t,W)
%
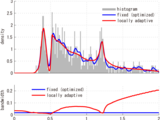
% Function `ssvkernel' returns an optimized kernel density estimate
% using a Gauss kernel function with bandwidths locally adapted to the data.
%
% Examples:
% >> x = 0.5-0.5*log(rand(1,1e3)); t = linspace(0,3,500);
% >> [y,t,optw] = ssvkernel(x,t);
% This example produces a vector of kernel density estimates, y, at points
% specified in a vector t, using locally adaptive bandwidths, optw
% (a standard deviation of a normal density function).
%
% >> ssvkernel(x);
% By calling the function without output arguments, the estimated density
% is displayed.
%
% Input arguments:
% x: Sample data vector.
% tin (optinal):
% Points at which estimation are computed.
% W (optinal):
% A vector of kernel bandwidths.
% If W is provided, the optimal bandwidth is selected from the
% elements of W.
% * Do not search bandwidths smaller than a sampling resolution of data.
% If W is not provided, the program searches the optimal bandwidth
% using a golden section search method.
%
% Output arguments:
% y: Estimated density
% t: Points at which estimation was computed.
% The same as tin if tin is provided.
% (If the sampling resolution of tin is smaller than the sampling
% resolution of the data, x, the estimation was done at smaller
% number of points than t. The results, t and y, are obtained by
% interpolating the low resolution sampling points.)
% optw: Optimal kernel bandwidth.
% gs: Stiffness constants of the variable bandwidth examined.
% The stifness constant is defined as a ratio of the optimal fixed
% bandwidth to a length of a local interval in which a fixed-kernel
% bandwidth optimization was performed.
% C: Cost functions of stiffness constants.
% conf95:
% Bootstrap confidence intervals.
% yb: Booststrap samples.
%
%
% Usage:
% >> [y,t,optw] = ssvkernel(x);
% When t is not given in the input arguments, i.e., the output argument t
% is generated automatically.
%
% >> W = linspace(0.01,1,20);
% >> [y,t,optw] = ssvkernel(x,t,W);
% The optimal bandwidth is selected from the elements of W.
%
% >> [y,t,optw,confb95,yb] = ssvkernel(x);
% This additionally computes 95% bootstrap confidence intervals, confb95.
% The bootstrap samples are provided as yb.
%
%
% Optimization principle:
% The optimization is based on a principle of minimizing
% expected L2 loss function between the kernel estimate and an unknown
% underlying density function. An assumption is merely that samples
% are drawn from the density independently each other.
%
% The locally adaptive bandwidth is obtained by iteratively computing
% optimal fixed-size bandwidths wihtihn local intervals. The optimal
% bandwidths are selected such that they are selected in the intervals
% that are \gamma times larger than the optimal bandwidths themselves.
% The paramter \gamma was optimized by minimizing the L2 risk estimate.
%
% The method is described in
% Hideaki Shimazaki and Shigeru Shinomoto
% Kernel Bandwidth Optimization in Spike Rate Estimation
% Journal of Computational Neuroscience 2010
% http://dx.doi.org/10.1007/s10827-009-0180-4
%
% * Instead of a Gaussian window described in the paper, a boxcar window is
% used in this program.
%
% For more information, please visit
% http://2000.jukuin.keio.ac.jp/shimazaki/res/kernel.html
%
% See also SSKERNEL, SSHIST
%
%
% Hideaki Shimazaki
% http://2000.jukuin.keio.ac.jp/shimazaki
Cite As
Hideaki Shimazaki (2026). ssvkernel(x,tin) (https://www.mathworks.com/matlabcentral/fileexchange/37374-ssvkernel-x-tin), MATLAB Central File Exchange. Retrieved .
General Information
- Version 1.3.0.0 (11.5 KB)
MATLAB Release Compatibility
- Compatible with any release
Platform Compatibility
- Windows
- macOS
- Linux
| Version | Published | Release Notes | Action |
|---|---|---|---|
| 1.3.0.0 | Bug fix: fixed a problem for large values
|
||
| 1.2.0.0 | fixed issues on compatibility to older versions of matlab. performance optimization. |
||
| 1.1.0.0 | correcting url in Description |
||
| 1.0.0.0 |