Example of how to use MATLAB Experiment Manager to test different classifiers for skin lesion classification using transfer learning.
https://github.com/ogemarques/matlab-experiment-manager-skin-lesion-classification
You are now following this Submission
- You will see updates in your followed content feed
- You may receive emails, depending on your communication preferences
Managing medical image classification experiments with MATLAB Experiment Manager App
This repository shows an example of how to use MATLAB Experiment Manager for a medical image classification task.
Experiment objective
To test the best combination of pretrained deep learning model and optimizer for a binary (malignant or benign) skin lesion classification task using transfer learning.
Common components
Dataset
Annotated images1 from the ISIC 2016 challenge, Task 3 (Lesion classification) dataset, consisting of 900 dermoscopic lesion images in JPEG format for training and validation, distributed in two classes (727 images were labeled as ‘benign’, 173 as ‘malignant’) plus 379 test images of the exact same format as the training data, and associated ground truth for all images.
Common hyperparameters
- Dataset partition (e.g., 70% for training, 30% for validation)
- Loss function
- Mini-batch size
- Initial learning rate
- Learning rate schedule
- Number of epochs
- Validation patience
- Validation frequency
- Performance metrics
Preprocessing scripts
- Image resizing (to the size expected by the input layer of each model)
- Image augmentation (e.g., translation, scaling and rotation)
Variable sets
A total of nine (3 x 3) combinations of:
- Model: ResNet-18, GoogLeNet or SqueezeNet
- Optimizer: ‘adam’, ‘sgdm’, or ‘rmsprop’
Requirements
- MATLAB 2020a or later
- Deep Learning Toolbox
- Deep Learning Toolbox™ Model for GoogLeNet Network support package
- Deep Learning Toolbox™ Model for ResNet-18 Network support package
- Deep Learning Toolbox™ Model for SqueezeNet Network support package
- Parallel Computing Toolbox (only required for training using a GPU)
Suggested steps
- Download or clone the repository.
- Open MATLAB.
- Edit the contents of the
dataFoldervariable in theexperiment1_setup.mlxfile to reflect the absolute path to your selected dataset2. - Open the Experiment Manager app (you can type
experimentmanagerin the MATLAB Command Window or select the app from the "APPS" ribbon). - Select the "New Project" option and click on the "Create" button.
- On the next screen, select the "Image Classification Using Transfer Learning" option and click on the "ADD" button.
- Specify a folder for your project.
- Edit the textbox containing the name of the Setup Function to reflect the name
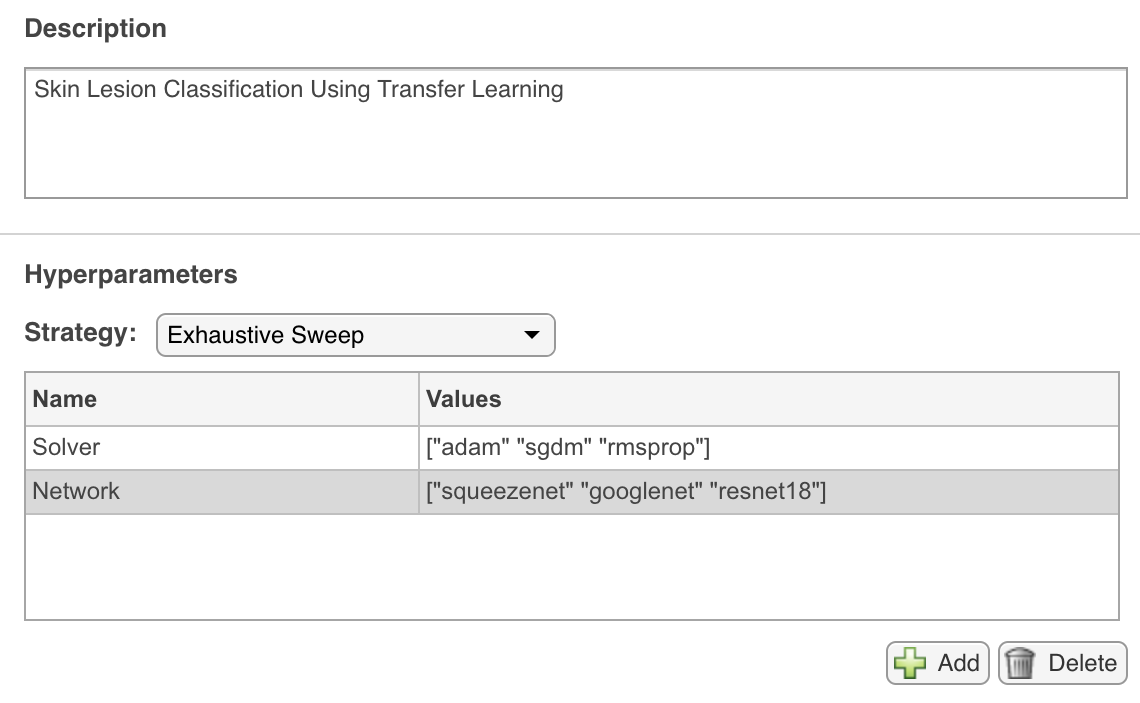
experiment1_setup. - Configure (i.e., edit manually) the hyperparameters table to reflect your choice of
Solver(["adam" "sgdm" "rmsprop]) andNetwork(["squeezenet" "googlenet" "resnet18"]) (see figure below).
- Copy the resulting
experiment1_setup.mlxfile to the project folder created by the Experiment Manager app. - (Optionally) click the "Use Parallel" button to run experiments in parallel.
- Click the "Run" button and watch the progress bars change as the different trials are run.
- Once the experiment concludes, explore/sort/filter/annotate the results.
- Save and close the project.
Additional remarks
- You are encouraged to expand and adapt the example to your needs.
- The choice of pretrained networks and their hyperparameters (learning rate, mini-batch size, number of epochs, etc.) is merely illustrative.
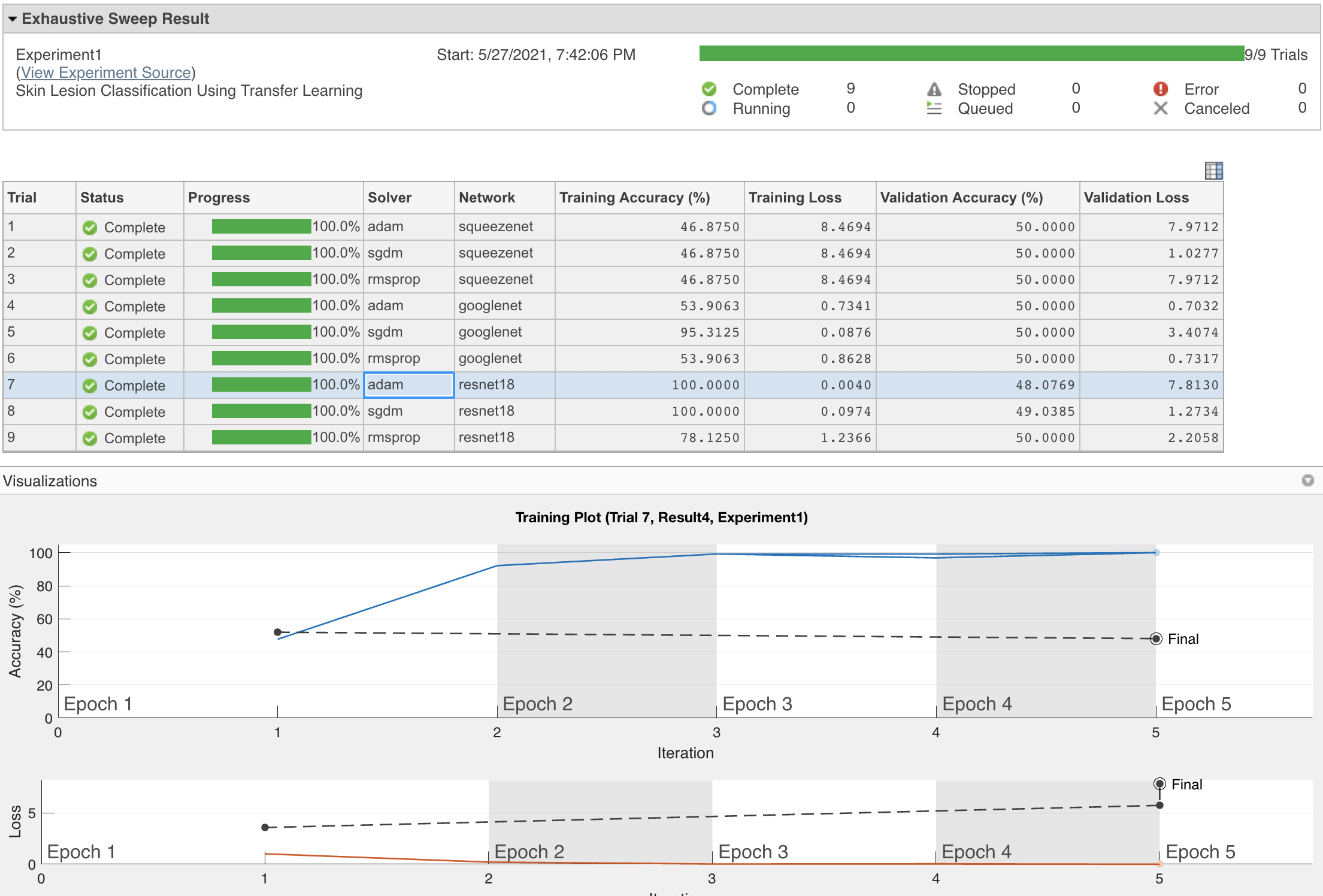
- Most of the resulting Network/Solver combinations do not show a stellar performance and all of them show signs of overfitting (see figure below).
- You are encouraged to (use Experiment Manager to) tweak those choices and find a better solution.
Notes
[1] This example uses a small subset of images to make it easier to get started without having to worry about large downloads and long training times.
[2] You can choose to use either the data folder or the balanced_data folder: both contain subfolders labeled benign and malignant.
Cite As
Oge Marques (2026). Managing medical image classification experiments (https://github.com/ogemarques/matlab-experiment-manager-skin-lesion-classification/releases/tag/1.0.4), GitHub. Retrieved .
General Information
- Version 1.0.4 (710 MB)
-
View License on GitHub
MATLAB Release Compatibility
- Compatible with any release
Platform Compatibility
- Windows
- macOS
- Linux
| Version | Published | Release Notes | Action |
|---|---|---|---|
| 1.0.4 | See release notes for this release on GitHub: https://github.com/ogemarques/matlab-experiment-manager-skin-lesion-classification/releases/tag/1.0.4 |
||
| 1.0.3 | Updated description |
||
| 1.0.2 | Placeholder |
||
| 1.0.1 | Updated main script |
||
| 1.0.0 |