mafdr
Estimate positive false discovery rate for multiple hypothesis testing
Syntax
Description
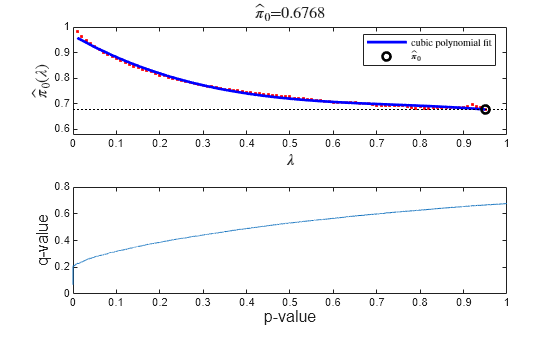
FDR = mafdr(PValues,Name,Value)'Showplot',true displays diagnostic plots of calculated results.
[
also returns FDR,Q,aPrioriProb] = mafdr(PValues,___)aPrioriProb, the estimated a priori
probability that the null hypothesis is true.
Examples
Input Arguments
Name-Value Arguments
Output Arguments
References
[1] Storey, John D. “A Direct Approach to False Discovery Rates.” Journal of the Royal Statistical Society: Series B (Statistical Methodology) 64, no. 3 (August 2002): 479–98.
[2] Benjamini, Y., and Hochberg, Y. 1995. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. Royal Stat. Soc. 57:289–300.
[3] Best, C.J.M., Gillespie, J.W., Yi, Y., Chandramouli, G.V.R., Perlmutter, M.A., Gathright, Y., Erickson, H.S., Georgevich, L., Tangrea, M.A., Duray, P.H., Gonzalez, S., Velasco, A., Linehan, W.M., Matusik, R.J., Price, D.K., Figg, W.D., Emmert-Buck, M.R., and Chuaqui, R.F. 2005. Molecular alterations in primary prostate cancer after androgen ablation therapy. Clin. Cancer Res. 11:6823–6831.
[4] Storey, J.D., and Tibshirani, R. 2003. Statistical significance for genomewide studies. Proc. Nat. Acad. Sci. 100:9440–9445.
[5] Storey, J.D., Taylor, J.E., and Siegmund, D. 2004. Strong control, conservative point estimation and simultaneous conservative consistency of false discovery rates: A unified approach. J. Royal Stat. Soc. 66:187–205.
Version History
Introduced in R2007a
See Also
mairplot | maloglog | mapcaplot | mattest | mavolcanoplot