mapcaplot
Create Principal Component Analysis (PCA) plot of microarray data
Description
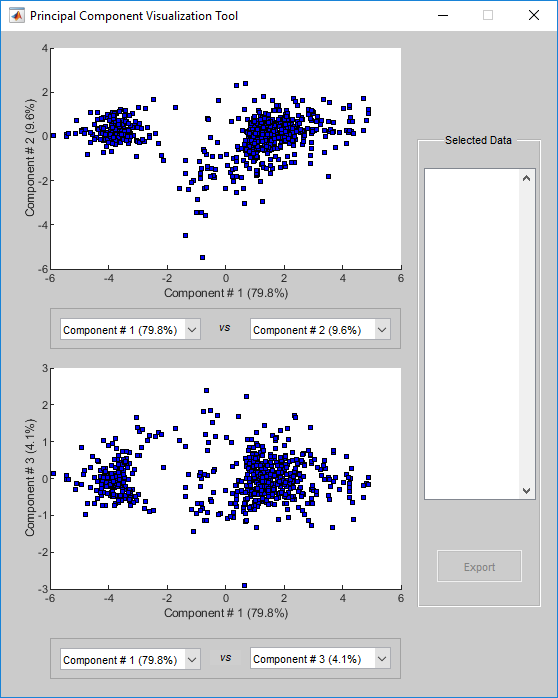
mapcaplot( creates 2-D scatter plots of
principal components of data)data. Once you plot the principal components,
you can:
Select principal components for the x and y axes from the drop-down list below each scatter plot.
Click a data point to display its label.
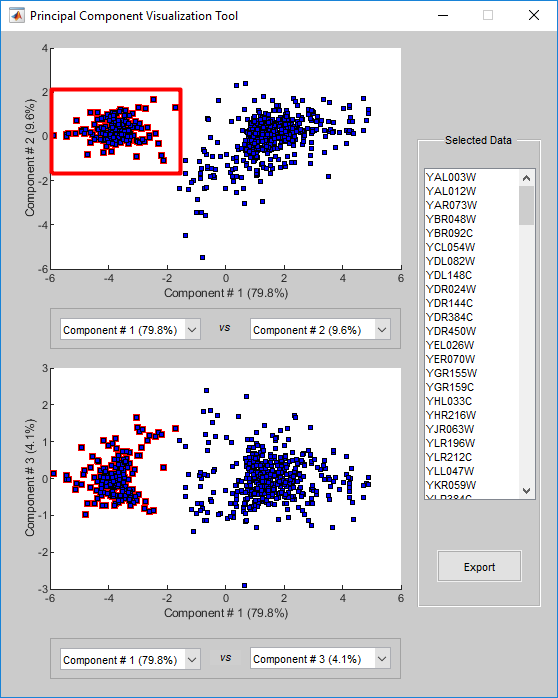
Select a subset of data points by dragging a box around them. Points in the selected region and the corresponding points in the other axes are then highlighted. The labels of the selected data points appear in the list box.
Select a label in the list box to highlight the corresponding data point in the plot. Press and hold Ctrl or Shift to select multiple data points.
Export the gene labels and indices to the MATLAB® workspace.
Examples
Input Arguments
References
[1] DeRisi, J.L., Iyer, V.R., and Brown, P.O. (1997). Exploring the metabolic and genetic control of gene expression on a genomic scale. Science 278, 680–686s.
Version History
Introduced before R2006a
See Also
clustergram | mattest | mavolcanoplot | pca