bwskel
Reduce all objects to lines in 2-D binary image or 3-D binary volume
Description
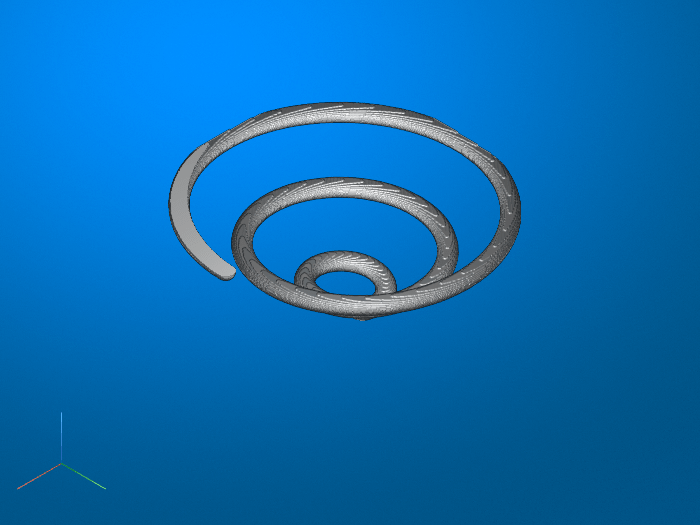
B = bwskel(A)A to 1-pixel wide
curved lines, without changing the essential structure of the image. This process,
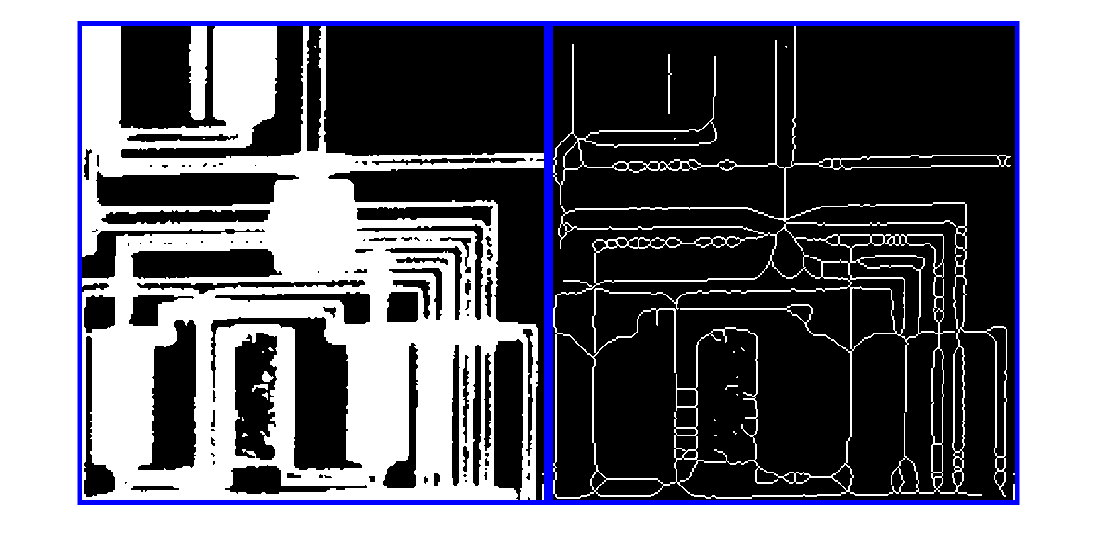
called skeletonization, extracts the centerline while
preserving the topology and Euler number (also known as the Euler characteristic) of
the objects.
Examples
Input Arguments
Output Arguments
Tips
While both
bwskelandbwmorphcan skeletonize 2-D images, the functions use different algorithms and can return different results. Thebwskelfunction uses 4-connectivity with 2-D images, while thebwmorphuses 8-connectivity. Thebwmorphoften creates a more accurate skeleton.bwskelassumes that foreground objects in the binary image are white (logicaltrue). If your image has a white background and black objects, then use the complement of your image as the input tobwskel. You can calculate the complement by using theimcomplementfunction.
Algorithms
The
bwskelfunction uses the medial axis transform.
References
[1] Ta-Chih Lee, Rangasami L. Kashyap and Chong-Nam Chu. Building skeleton models via 3-D medial surface/axis thinning algorithms. Computer Vision, Graphics, and Image Processing, 56(6):462-478, 1994.
[2] Kerschnitzki, M, Kollmannsberger, P, Burghammer, M. et al. Architecture of the osteocyte network correlates with bone material quality. Journal of Bone and Mineral Research, 28(8):1837-1845, 2013.