uniquetol
Unique values within tolerance
Syntax
Description
C = uniquetol(A,tol)A using tolerance tol.
Two values, u and v, are within
tolerance if
abs(u-v) <= tol*max(abs(A(:)))
That is, uniquetol scales tol input based on the
magnitude of the data.
uniquetol is similar to unique.
unique performs exact comparisons, while
uniquetol performs comparisons using a
tolerance.
C = uniquetol(A,___,occurrence)occurrence for either of the previous syntaxes.
[___] = uniquetol(___,
specifies additional parameters for determining the unique elements using one or
more name-value arguments. For example,
Name=Value)uniquetol(A,ByRows=true) determines the unique rows in
A.
Examples
Create a vector x. Obtain a second vector y by transforming and untransforming x. This transformation introduces round-off differences in y.
x = (1:6)'*pi; y = 10.^log10(x);
Verify that x and y are not identical by taking the difference.
x-y
ans = 6×1
10-14 ×
0.0444
0
0
0
0
-0.3553
Concatenate the vectors x and y. Then, use unique to find the unique elements. The unique function performs exact comparisons and determines that some values in x are not exactly equal to values in y. These are the same elements that have a nonzero difference in x-y. Thus, C contains values that appear to be duplicates.
A = [x; y]; C = unique(A)
C = 8×1
3.1416
3.1416
6.2832
9.4248
12.5664
15.7080
18.8496
18.8496
Use uniquetol to perform the comparison using a small tolerance. uniquetol treats elements that are within tolerance as equal.
Ctol = uniquetol(A)
Ctol = 6×1
3.1416
6.2832
9.4248
12.5664
15.7080
18.8496
By default, uniquetol looks for unique elements that are within tolerance, but it also can find unique rows of a matrix that are within tolerance.
Create a numeric matrix, x. Obtain a second matrix, y, by transforming and untransforming x. This transformation introduces round-off differences to y.
x = [0.05 0.11 0.18; 0.18 0.21 0.29; 0.34 0.36 0.41; 0.46 0.52 0.76]; y = log10(10.^x);
Concatenate the matrices x and y. Then, use unique to find the unique rows. The unique function performs exact comparisons and determines that all of the rows in the concatenated matrix [x; y] are unique, even though some of the rows differ by only a small amount.
A = [x; y];
C = unique(A,"rows")C = 8×3
0.0500 0.1100 0.1800
0.0500 0.1100 0.1800
0.1800 0.2100 0.2900
0.1800 0.2100 0.2900
0.3400 0.3600 0.4100
0.3400 0.3600 0.4100
0.4600 0.5200 0.7600
0.4600 0.5200 0.7600
Use uniquetol to find the unique rows. uniquetol treats rows that are within tolerance as equal.
Ctol = uniquetol(A,ByRows=true)
Ctol = 4×3
0.0500 0.1100 0.1800
0.1800 0.2100 0.2900
0.3400 0.3600 0.4100
0.4600 0.5200 0.7600
Create a vector, x. Obtain a second vector, y, by transforming and untransforming x. This transformation introduces round-off differences to some elements in y.
x = (1:5)'*pi; y = 10.^log10(x);
Concatenate the vectors x and y. Then, use uniquetol to reconstruct A, treating the values that are within tolerance as equal.
A = [x; y]; [C,IA,IC] = uniquetol(A); newA = C(IC)
newA = 10×1
3.1416
6.2832
9.4248
12.5664
15.7080
3.1416
6.2832
9.4248
12.5664
15.7080
You can use newA with == or functions that use exact equality like isequal or unique in subsequent code.
D1 = unique(A)
D1 = 6×1
3.1416
3.1416
6.2832
9.4248
12.5664
15.7080
D2 = unique(newA)
D2 = 5×1
3.1416
6.2832
9.4248
12.5664
15.7080
Control which elements uniquetol selects as being unique by specifying the occurrence option.
Create a vector and find which elements are unique within a tolerance of 1e-1.
A = [1 1.1 1.11 1.12 1.13 2]; C = uniquetol(A,1e-1)
C = 1×2
1 2
Since the first five elements in A all have similar values with respect to the tolerance of 1e-1, only the lowest value among them is selected as being unique. This is because uniquetol begins with the lowest value in a and does not find a new element that is not within tolerance until the 2 at the end of the vector.
Use the "highest" option to specify that uniquetol should begin with the highest value in A. Now, the 1.13 element is selected as being unique since uniquetol works down from the highest values.
C2 = uniquetol(A,1e-1,"highest")C2 = 1×2
1.1300 2.0000
Create a cloud of 2-D sample points constrained to be inside a circle of radius 0.5 centered at the point .
x = rand(10000,2); insideCircle = sqrt((x(:,1)-.5).^2+(x(:,2)-.5).^2)<0.5; A = x(insideCircle,:);
Find a reduced set of points, such that each point of the original dataset is within tolerance of a point.
tol = 0.05; C = uniquetol(A,tol,ByRows=true);
Plot the original data set and the reduced set of points. All of the points in the reduced set are members of the original data set and are at least a distance tol apart.
plot(A(:,1),A(:,2),"c.") hold on axis equal plot(C(:,1),C(:,2),"r.",MarkerSize=10) legend("Original Data","Reduced Data")
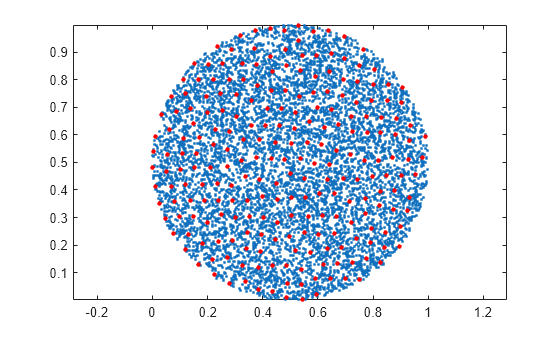
Create a vector of random numbers and determine the unique elements using a tolerance. Specify OutputAllIndices as true to return all of the indices for the elements that are within tolerance of the unique values.
rng default
A = rand(100,1);
[C,IA] = uniquetol(A,1e-2,OutputAllIndices=true);Find the average value of the elements that are within tolerance of the value C(2).
C(2)
ans = 0.0318
allA = A(IA{2})allA = 3×1
0.0357
0.0318
0.0344
aveA = mean(allA)
aveA = 0.0340
By default, uniquetol uses a tolerance test of the form abs(u-v) <= tol*DS, where DS automatically scales based on the magnitude of the input data. You can specify a different DS value to use with the DataScale name-value argument. However, absolute tolerances (where DS is a scalar) do not scale based on the magnitude of the input data.
First, compare two small values that are a distance eps apart. Specify tol and DS to make the within tolerance equation: abs(u-v) <= 10^-6.
x = 0.1; uniquetol([x exp(log(x))],10^-6,DataScale=1)
ans = 0.1000
Next, increase the magnitude of the values. The round-off error in the calculation exp(log(x)) is proportional to the magnitude of the values, specifically to eps(x). Even though the two large values are a distance eps from one another, eps(x) is now much larger. Therefore, 10^-6 is no longer a suitable tolerance.
x = 10^10; uniquetol([x exp(log(x))],10^-6,DataScale=1)
ans = 1×2
1010 ×
1.0000 1.0000
Correct this issue by using the default (scaled) value of DS.
format long
Y = [0.1 10^10];
uniquetol([Y exp(log(Y))])ans = 1×2
1010 ×
0.000000000010000 1.000000000000000
Create a set of random 2-D points, then use uniquetol to group the points into vertical bands that have a similar (within tolerance) x-coordinate. Use these options with uniquetol:
Specify
ByRowsastruesince the point coordinates are in the rows ofA.Specify
OutputAllIndicesastrueto return the indices for all points that have an x-coordinate within tolerance of each other.Specify
DataScaleas[1 Inf]to use an absolute tolerance for thex-coordinate while ignoring they-coordinate.
A = rand(1000,2); DS = [1 Inf]; [C,IA] = uniquetol(A,0.1,ByRows=true,OutputAllIndices=true,DataScale=DS);
Plot the points and average value for each band.
hold on for k = 1:length(IA) plot(A(IA{k},1),A(IA{k},2),".") meanAi = mean(A(IA{k},:)); plot(meanAi(1),meanAi(2),"xr") end
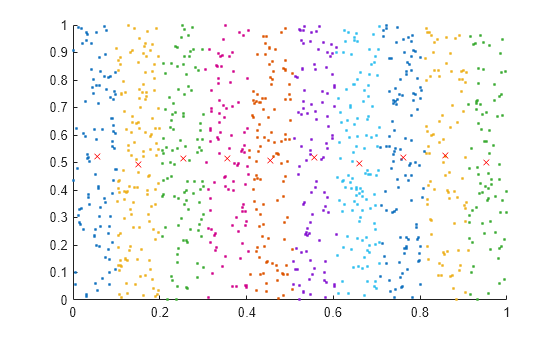
Input Arguments
Query array, specified as a real and full array.
Data Types: single | double
Comparison tolerance, specified as a positive real scalar.
uniquetol scales the tol input
using the maximum absolute value in input array A. Then
uniquetol uses the resulting scaled comparison
tolerance to determine which elements in A are unique. If
two elements in A are within tolerance of each other,
then uniquetol considers them to be equal.
Two values, u and v, are within
tolerance if abs(u-v) <= tol*max(abs(A)).
To specify an absolute tolerance, specify both tol and
the DataScale name-value argument.
Example: 0.05
Example: 1e-8
Example: eps
Data Types: single | double
Occurrence of unique values, specified as one of the options in this
table. The value of occurrence determines which elements
uniquetol selects as being unique.
| Option | Description |
|---|---|
"lowest" |
|
"highest" |
|
Example: C = uniquetol(A,tol,"highest")
Example: C = uniquetol([1 2 3],2,"highest") returns
3, since uniquetol starts with
the highest value in the input and all other values are within
tolerance.
Data Types: string | char
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: C = uniquetol(A,ByRows=true)
Whether to return all indices for repeated elements, specified as
logical 0 (false) or
1 (true).
When OutputAllIndices is false,
IA is a column vector that contains indices to
the first occurrence of repeated elements.
When OutputAllIndices is true,
IA is a cell array that contains the indices for
all elements in A that are within tolerance of a
value in C. Each cell in IA
corresponds to a value in C, and the values in each
cell correspond to locations in A.
Example: [C,IA] =
uniquetol(A,tol,OutputAllIndices=true)
Whether to compare the rows of A, specified as
logical 0 (false) or
1 (true). Specify
ByRows to find rows in A that
are unique, within tolerance.
When ByRows is true:
Amust be a 2-D array.uniquetolcompares the rows ofAby considering each column separately. For two rows to be within tolerance of one another, each column has to be in tolerance.Each row in
Ais within tolerance of a row inC. However, no two rows inCare within tolerance of each other.
Two rows, u and v, are within
tolerance if all(abs(u-v) <=
tol*max(abs(A),[],1)).
Example: C =
uniquetol(A,tol,ByRows=true)
Scale of data, specified as a scalar or vector. Specify DataScale as a
numeric scalar, DS, to change the tolerance test to
be abs(u-v) <= tol*DS.
When you specify both DataScale and ByRows,
DataScale can be a vector. In this case, each
element of the vector specifies DS for a
corresponding column in A. If a value in the
DataScale vector is Inf, then
uniquetol ignores the corresponding column in
A.
Example: C = uniquetol(A,DataScale=1)
Example: [C,IA,IC] = uniquetol(A,ByRows=true,DataScale=[eps(1) eps(10)
eps(100)])
Data Types: single | double
Whether to preserve the range of A, specified as
logical 0 (false) or
1 (true). If
PreserveRange is true, then
the minimum and maximum values in the output C are
the same as those in A.
If PreserveRange is false and
the minimum and maximum values in A are within the
tolerance tol of each other, then
uniquetol returns only one of the
values.
Example: C =
uniquetol(A,tol,PreserveRange=true)
Output Arguments
Unique elements within tolerance, returned as a vector or matrix.
If A is a row vector, then C is also
a row vector. Otherwise, C is a column vector. The
elements in C are sorted in ascending order. Each element
in A is within tolerance of an element in
C, but no two elements in C are
within tolerance of each other.
If ByRows is true, then
C is a matrix containing the unique rows in
A. In this case, the rows in C are
sorted in ascending order by the first column. Each row in
A is within tolerance of a row in
C, but no two rows in C are within
tolerance of each other.
Index to A, returned as a column vector of indices to the first occurrence
of repeated elements or as a cell array. IA generally
satisfies C = A(IA), with these exceptions:
If the
ByRowsoption istrue, thenC = A(IA,:).If
OutputAllIndicesistrue, thenIAis a cell array andC(i)andA(IA{i})are within tolerance of each other.
Index to C, returned as a column vector of
indices. IC satisfies the following properties,
where ~ means the values are within tolerance of
each other.
If
Ais a vector, thenA~C(IC).If
Ais a matrix, thenA(:)~C(IC).If
ByRowsistrue, thenA~C(IC,:).
Algorithms
uniquetol sorts the input lexicographically, and then starts at
the lowest or highest value to find unique values within tolerance. As a result,
changing the sorting of the input could change the output. For example,
uniquetol(-A) might not give the same results as
-uniquetol(A).
There can be multiple valid C outputs that satisfy the condition,
no two elements in C are within tolerance of each
other. The uniquetol function can return several of
the valid outputs, depending on whether the value of occurrence is
"highest" or "lowest" and whether
PreserveRange is specified.
Extended Capabilities
Usage notes and limitations:
The
OutputAllIndicesname-value argument is not supported.The query array must be a vector unless the value of
ByRowsistrue.
Refer to the usage notes and limitations in the C/C++ Code Generation section. The same usage notes and limitations apply to GPU code generation.
The uniquetol function fully supports
thread-based environments. For more information, see Run MATLAB Functions in Thread-Based Environment.
The uniquetol function
supports GPU array input with these usage notes and limitations:
The
occurrenceargument for occurrence of unique values is not supported.The
ByRows,OutputAllIndices, andPreserveRangename-value arguments are not supported.64-bit integers are not supported.
For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2015aYou can generate C/C++ code for the uniquetol
function.
The occurrence argument controls whether the algorithm begins
with the highest or lowest elements in the input data. The
PreserveRange name-value argument specifies whether the range
of the output data should be the same as the input data.
See Also
unique | isapprox | ismembertol | ismember | eps
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)